PathAsst: A Generative Foundation AI Assistant Towards Artificial General Intelligence of Pathology
PathAsst: 迈向病理学通用人工智能 (AGI) 的生成式基础 AI 助手
Yuxuan $\mathbf{Sun}^{1,2,* }$ , Chenglu Zhu2,* , Sunyi Zheng2, Kai Zhang3, Lin Sun4, Zhongyi Shui1,2, Yunlong Zhang1,2, Honglin Li1,2, Lin Yang2,
Yuxuan $\mathbf{Sun}^{1,2,* }$、Chenglu Zhu2,* 、Sunyi Zheng2、Kai Zhang3、Lin Sun4、Zhongyi Shui1,2、Yunlong Zhang1,2、Honglin Li1,2、Lin Yang2
1College of Computer Science and Technology, Zhejiang University, China 2Research Center for Industries of the Future and School of Engineering, Westlake University, China 3 Department of Computer Science and Engineering, The Ohio State University, USA 4School of Computer and Computing Science, Hangzhou City University, China {sunyuxuan,yanglin} $@$ westlake.edu.cn
1浙江大学计算机科学与技术学院,中国 2西湖大学未来产业研究中心及工学院,中国 3俄亥俄州立大学计算机科学与工程系,美国 4杭州城市学院计算机与计算科学学院,中国 {sunyuxuan,yanglin} $@$ westlake.edu.cn
Abstract
摘要
As advances in large language models (LLMs) and multimodal techniques continue to mature, the development of general-purpose multimodal large language models (MLLMs) has surged, offering significant applications in interpreting natural images. However, the field of pathology has largely remained untapped, particularly in gathering high-quality data and designing comprehensive model frameworks. To bridge the gap in pathology MLLMs, we present PathAsst, a multimodal generative foundation AI assistant to revolutionize diagnostic and predictive analytics in pathology. The development of PathAsst involves three pivotal steps: data acquisition, CLIP model adaptation, and the training of PathAsst’s multimodal generative capabilities. Firstly, we collect over 207K high-quality pathology image-text pairs from authoritative sources. Leveraging the advanced power of ChatGPT, we generate over 180K instruction-following samples. Furthermore, we devise additional instruction-following data specifically tailored for invoking eight pathology-specific sub-models we prepared, allowing the PathAsst to effectively collaborate with these models, enhancing its diagnostic ability. Secondly, by leveraging the collected data, we construct PathCLIP, a pathology-dedicated CLIP, to enhance PathAsst’s capabilities in interpreting pathology images. Finally, we integrate PathCLIP with the Vicuna-13b and utilize pathology-specific instruction-tuning data to enhance the multimodal generation capacity of PathAsst and bolster its synergistic interactions with sub-models. The experimental results of PathAsst show the potential of harnessing AI-powered generative foundation model to improve pathology diagnosis and treatment pro- cesses. We open-source our dataset, as well as a comprehensive toolkit for extensive pathology data collection and preprocessing at https://github.com/super james sy x/GenerativeFoundation-AI-Assistant-for-Pathology.
随着大语言模型(LLM)和多模态技术的持续成熟,通用多模态大语言模型(MLLM)的发展呈现出爆发式增长,在自然图像解读领域展现出重要应用价值。然而病理学领域仍存在大量未开发潜力,特别是在高质量数据收集和综合模型框架设计方面。为填补病理学MLLM的空白,我们推出PathAsst——一个革新病理学诊断与预测分析的多模态生成式基础(Generative Foundation)AI助手。PathAsst的开发包含三个关键步骤:数据采集、CLIP模型适配,以及PathAsst多模态生成能力的训练。
首先,我们从权威来源收集了超过207K组高质量病理图文配对数据。借助ChatGPT的先进能力,我们生成了超过180K条指令跟随样本。此外,我们还专门设计了用于调用八个预置病理学子模型的指令数据,使PathAsst能有效协同这些模型以增强诊断能力。
其次,基于收集的数据,我们构建了病理学专用CLIP模型PathCLIP,以提升PathAsst解析病理图像的能力。最后,我们将PathCLIP与Vicuna-13b集成,并利用病理学专用指令调优数据来增强PathAsst的多模态生成能力,强化其与子模型的协同交互。
PathAsst的实验结果表明,利用AI驱动的生成式基础模型改进病理诊疗流程具有巨大潜力。我们在https://github.com/superjames sy x/GenerativeFoundation-AI-Assistant-for-Pathology开源了数据集,以及用于大规模病理数据收集与预处理的完整工具包。
Introduction
引言
In recent years, artificial intelligence has made remarkable strides across various fields (Liu et al. 2022b; Zhuang et al. 2021). This is particularly evident in pathology, which has undergone a profound transformation with the introduction of digital pathology and advanced deep learning techniques. The increasing availability of digitized his to pathology data, coupled with the exponential growth in the size and complexity of pathology datasets, has necessitated the development of more sophisticated tools to enhance the analytical efficiency of pathologists.
近年来,人工智能在各个领域取得了显著进展 (Liu et al. 2022b; Zhuang et al. 2021)。这一趋势在病理学领域尤为明显,随着数字病理学 (digital pathology) 和先进深度学习技术的引入,该学科经历了深刻变革。数字化病理数据的日益普及,加之病理数据集规模和复杂性的指数级增长,亟需开发更先进的工具来提升病理学家的分析效率。
Simultaneously, there has been an upsurge interest in LLMs, with numerous researchers focusing on their development and application. The ultimate goal is to create models with general artificial intelligence capabilities. Among the most prominent examples are OpenAI’s ChatGPT and GPT-4. These models have showcased impressive capabilities in human interaction by training through instruction tuning and human feedback, thereby fueling the community’s enthusiasm for LLMs.
与此同时,大语言模型(LLM)的研究热潮兴起,众多学者聚焦于其开发与应用,终极目标是构建具备通用人工智能(AGI)能力的模型。其中最典型的代表当属OpenAI的ChatGPT和GPT-4。这些模型通过指令微调(instruction tuning)和人类反馈训练,在人机交互中展现出惊人能力,从而激发了学界对大语言模型的研发热情。
In the open-source community, LLaMA (Touvron et al. 2023) has emerged as a compelling model that exhibits performance on par with GPT-3 (Brown et al. 2020), provid- ing promising opportunities for further development. Subsequent models, such as Alpaca (Taori et al. 2023) and Vicuna (Chiang et al. 2023), take advantage of LLaMA and leverage the instruction tuning techniques, enabling them even outperform ChatGPT in certain tasks. Researchers have also explored the realm of multimodal models, creating innovative approaches such as LLaVA (Liu et al. 2023a) and MiniGPT-4 (Zhu et al. 2023). These models demonstrate impressive capabilities in comprehending and interpreting multimodal data, showcasing the advancements in the field.
在开源社区中,LLaMA (Touvron et al. 2023) 已成为一个引人注目的模型,其性能与 GPT-3 (Brown et al. 2020) 相当,为后续发展提供了广阔空间。后续模型如 Alpaca (Taori et al. 2023) 和 Vicuna (Chiang et al. 2023) 基于 LLaMA 并采用指令微调技术,使其在某些任务中甚至超越 ChatGPT。研究者们还探索了多模态模型领域,开发出 LLaVA (Liu et al. 2023a) 和 MiniGPT-4 (Zhu et al. 2023) 等创新方案,这些模型在理解和解析多模态数据方面展现出卓越能力,彰显了该领域的重大进展。
However, while these advanced MLLMs primarily focus on natural images, the field of pathology faces a notable gap due to the scarcity of high-quality data and limited exploration of model frameworks, which results in a deficiency of pathology-specific MLLMs. In this study, we aim to bridge this gap by exploring both high-quality pathology data collection and the potential application of MLLMs within the pathology domain. We outline our contributions as follows:
然而,尽管这些先进的多模态大语言模型(MLLM)主要聚焦于自然图像领域,病理学领域却因高质量数据稀缺和模型框架探索有限而存在显著空白,导致缺乏针对病理学的专用MLLM。本研究旨在通过探索高质量病理数据收集及MLLM在病理学领域的潜在应用来弥合这一缺口。我们的贡献概述如下:
• We gather diverse pathology image-caption pairs from authoritative sources. Through a meticulous process of data cleaning and optimization, we create the PathCap dataset, comprising 207K high-quality samples. • We introduce PathCLIP, a pathology-specific CLIP model trained on the PathCap. Compared to prior models, PathCLIP shows superior proficiency in understanding pathology data, achieving state-of-the-art results in pathology image retrieval and zero-shot classification.
• 我们从权威来源收集多样化的病理学图像-标题对。通过严格的数据清洗和优化流程,构建了包含20.7万高质量样本的PathCap数据集。
• 我们提出PathCLIP——基于PathCap训练的病理学专用CLIP模型。相较于现有模型,PathCLIP在病理学数据理解方面展现出卓越能力,在病理图像检索和零样本分类任务中达到最先进性能。
• We integrate PathCLIP and Vicuna-13b to develop PathAsst, a multimodal generative foundational model tailored for pathology. Utilizing the PathCap dataset, we prompt ChatGPT to generate the Path Instruct dataset, which consists of 180K pathology multimodal instruction-following samples. These samples are employed to train PathAsst’s generative capabilities. Additionally, we prepare eight pathology-specific sub-models, supplemented with instruction-following data for various scenarios that necessitate sub-model invocation. This equips PathAsst with the ability to discern when to utilize these models for optimal results.
• 我们将 PathCLIP 与 Vicuna-13b 集成,开发出专为病理学设计的跨模态生成式基础模型 PathAsst。通过 PathCap 数据集,我们提示 ChatGPT 生成包含 18 万例病理学多模态指令跟随样本的 Path Instruct 数据集,用于训练 PathAsst 的生成能力。此外,我们构建了八个病理学子模型,并针对需要调用子模型的不同场景补充指令跟随数据,使 PathAsst 能够智能判断何时调用这些模型以获得最佳结果。
Related Work
相关工作
Large Language Model (LLM). In the early stages, breakthrough models like BERT (Devlin et al. 2018) and GPT (Radford et al. 2018), were introduced, drawing inspiration from the transformer architecture. These models ignited significant interest in the natural language processing (NLP) domain and signaled the beginning of large-scale models in this field. Initially, the full potential of generative models remained largely unexplored. However, in recent years, as the generative model continue to scale up, more powerful models such as GPT-3 (Brown et al. 2020), T5 (Raffel et al. 2020), PaLM (Chowdhery et al. 2022), and OPT (Zhang et al. 2022) are developed. Their emergent abilities (Wei et al. 2022) lead these larger models to display markedly superior performance on complex tasks compared to their smaller counterparts. Furthermore, the introduction of instruction tuning techniques (Ouyang et al. 2022; Wang et al. 2022b,a), specifically in the realm of LLM, enables the generation of more controllable, practical, and task-specific results. This revolutionary enhancement significantly boosts the zero-shot learning abilities of large models, as exemplified by Instruct GP T (Ouyang et al. 2022), GPT-4 (OpenAI 2023), FLAN-T5 (Chung et al. 2022), and FLAN-PaLM (Chung et al. 2022).
大语言模型 (LLM)。早期阶段,受 Transformer 架构启发,BERT (Devlin et al. 2018) 和 GPT (Radford et al. 2018) 等突破性模型相继问世。这些模型点燃了自然语言处理 (NLP) 领域的广泛关注,标志着该领域大规模模型时代的开端。最初,生成式模型的全部潜力尚未得到充分发掘。但近年来,随着生成式模型规模持续扩大,更强大的模型如 GPT-3 (Brown et al. 2020)、T5 (Raffel et al. 2020)、PaLM (Chowdhery et al. 2022) 和 OPT (Zhang et al. 2022) 相继涌现。它们的涌现能力 (Wei et al. 2022) 使这些大模型在复杂任务上展现出远超小模型的性能。此外,指令微调技术 (Ouyang et al. 2022; Wang et al. 2022b,a) 的引入,特别是针对大语言模型的技术,使得模型能够生成更可控、实用且任务专属的结果。这一革命性提升显著增强了大模型的零样本学习能力,典型代表包括 Instruct GPT (Ouyang et al. 2022)、GPT-4 (OpenAI 2023)、FLAN-T5 (Chung et al. 2022) 和 FLAN-PaLM (Chung et al. 2022)。
Multimodal Large Language Model (MLLM). Recent advancements in large-scale multimodal models can be primarily divided into two branches. The first branch is developed based on the LangChain (Chase 2022) approach, where LLM collaborates with various specialized visual models to generate results. Prominent representatives of this branch include Visual ChatGPT (Wu et al. 2023) and MM-REACT (Yang et al. 2023). The second branch is implemented by integrating the feature outputs from visual models into the token sequence inputs of the LLM, enabling multimodal generation. This method is represented in models such as BLIP-2 (Li et al. 2023), PaLME (Driess et al. 2023) and Flamingo (Alayrac et al. 2022). Building upon the instruction-tuning techniques inspired by the LLM community, researchers create multimodal instruction-following datasets to perform MLLM training. This approach promptes the development of models such as LLaVA (Liu et al. 2023a), MiniGPT-4 (Zhu et al. 2023) and LLaMA-Adapter V2 (Gao et al. 2023). These models demonstrate impressive performance in solving multimodal tasks, as well as advanced multimodal chat capabilities.
多模态大语言模型 (Multimodal Large Language Model, MLLM) 。当前大规模多模态模型的发展主要分为两个方向。第一个方向基于 LangChain (Chase 2022) 框架实现,通过大语言模型与各类专业视觉模型协作生成结果,其典型代表包括 Visual ChatGPT (Wu et al. 2023) 和 MM-REACT (Yang et al. 2023) 。第二个方向通过将视觉模型的特征输出整合至大语言模型的token输入序列,实现多模态生成能力,该方法在 BLIP-2 (Li et al. 2023) 、PaLME (Driess et al. 2023) 和 Flamingo (Alayrac et al. 2022) 等模型中得以体现。受大语言模型社区指令微调技术的启发,研究者通过构建多模态指令跟随数据集进行 MLLM 训练,由此催生了 LLaVA (Liu et al. 2023a) 、MiniGPT-4 (Zhu et al. 2023) 和 LLaMA-Adapter V2 (Gao et al. 2023) 等模型。这些模型不仅在解决多模态任务时表现卓越,还展现出先进的多模态对话能力。
Multimodal Model for Pathology. While there are numerous applications for multimodal models in natural image analysis, their use in pathological image analysis has been relatively limited to date. The majority of methods employ approaches that combine vision encoder with LSTM (Liu et al. 2023b; Zhang et al. 2019a,b), yielding fairly satisfactory results. TraP-VQA (Naseem, Khushi, and Kim 2022) is the first attempt to employ vision-language transformer in pathology image processing, which is tested on the PathVQA dataset (He et al. 2020) to generate interpret able answers. More recently, Huang et al. (Huang et al. 2023) compile a large-scale dataset of pathology image-text pairs, sourced from social media platforms such as Twitter. They utilize contrastive vision-language pre training to establish a foundational model for pathology, demonstrating promising results in pathology zero-shot image-text cross-modal retrieval and zero-shot image classification.
病理学多模态模型。尽管多模态模型在自然图像分析中有众多应用,但迄今为止其在病理图像分析中的使用仍相对有限。大多数方法采用视觉编码器与LSTM结合的方案 (Liu et al. 2023b; Zhang et al. 2019a,b),取得了较为满意的效果。TraP-VQA (Naseem, Khushi, and Kim 2022) 首次尝试在病理图像处理中应用视觉语言Transformer,该模型在PathVQA数据集 (He et al. 2020) 上测试以生成可解释的答案。最近,Huang等人 (Huang et al. 2023) 从Twitter等社交媒体平台收集构建了大规模病理图文配对数据集,通过对比式视觉语言预训练建立了病理学基础模型,在病理零样本图文跨模态检索和零样本图像分类任务中展现了优异性能。
Multimodal Datasets. Numerous researchers have been dedicating their efforts to contribute valuable datasets that facilitate the advancement of models in the aforementioned domains. For instance, in the general domain, the community has successfully constructed various datasets, such as CC (Changpinyo et al. 2021) and LAION (Schuhmann et al. 2022). In the biomedical field, researchers have released datasets like ROCO (Pelka et al. 2018), MedICAT (Subramanian et al. 2020), and PMC-OA (Lin et al. 2023). In the pathology domain, researchers have recently built the OpenPath (Huang et al. 2023) dataset by crawling Twitter.
多模态数据集。众多研究者致力于贡献有价值的数据集,以推动上述领域模型的进步。例如,在通用领域,社区已成功构建了多种数据集,如CC (Changpinyo et al. 2021) 和 LAION (Schuhmann et al. 2022) 。在生物医学领域,研究者发布了ROCO (Pelka et al. 2018) 、MedICAT (Subramanian et al. 2020) 和 PMC-OA (Lin et al. 2023) 等数据集。在病理学领域,研究者近期通过爬取Twitter构建了OpenPath (Huang et al. 2023) 数据集。
Despite significant progress in the field, the domain of MLLM specifically adapted for pathology remains largely untapped. Current models, primarily designed for caption generation, often under perform when compared to specialized professional pathology models. Furthermore, regarding pathology MLLM dataset construction, existing datasets such as ROCO, MedICAT, and PMC-OA are not specifically tailored for this field. The only large-scale dataset, OpenPath, primarily sources its data from Twitter, where the image-text correlation is relatively weak, thus posing challenges for MLLM training. Moreover, the image-text pairs in OpenPath require access to the Twitter API, which carries a significant cost. As a result, there is still a substantial lack of high-quality image-caption datasets in the field of pathology. To bridge this gap, we develop two comprehensive pathology multimodal datasets. Building on these datasets, we utilize the power of instruction tuning to significantly improve MLLM’s capability in interpreting pathology images.
尽管该领域取得了显著进展,但专门针对病理学优化的多模态大语言模型(MLLM)方向仍存在大量空白。当前主要面向描述生成设计的模型,其性能往往逊色于专业病理学模型。在病理学MLLM数据集构建方面,现有ROCO、MedICAT和PMC-OA等数据集并非为该领域专门设计。唯一的大规模数据集OpenPath主要从Twitter获取数据,其图文关联性较弱,这为MLLM训练带来了挑战。此外,OpenPath中的图文对需通过Twitter API获取,成本较高。因此,病理学领域仍严重缺乏高质量的图像-描述数据集。为填补这一空白,我们开发了两个综合性病理学多模态数据集。基于这些数据集,我们利用指令调优技术显著提升了MLLM解析病理学图像的能力。
Pathology Dataset Construction
病理学数据集构建
In this paper, we propose two datasets tailored for pathology: PathCap and Path Instruct. The PathCap contains 207K high-quality pathology image-caption pairs. Among them, 197K are collected from PubMed and internal pathology guidelines books, while an additional 10K annotations are provided by expert cytologists specializing in liquid-based cytology (LBC). The Path Instruct dataset consists of 180K samples and includes two parts of instruction-following data. The first part is generated by prompting ChatGPT based on curated pathology image-text pairs (refer to step 4 in the subsequent data processing introduction). The second section includes multimodal instruction-following data tailored for model invocation, ensuring the effective use of specialized pathology models based on user intent and image features.
本文提出了两个专为病理学定制的数据集:PathCap和Path Instruct。PathCap包含207K个高质量病理图像-标题对,其中197K来自PubMed和内部病理指南书籍,另有10K标注由专注于液基细胞学(LBC)的专家细胞学家提供。Path Instruct数据集包含180K样本,由两部分指令遵循数据组成:第一部分通过基于精选病理图像-文本对提示ChatGPT生成(参见后续数据处理介绍中的步骤4),第二部分包含为模型调用定制的多模态指令遵循数据,确保根据用户意图和图像特征有效调用专业病理模型。
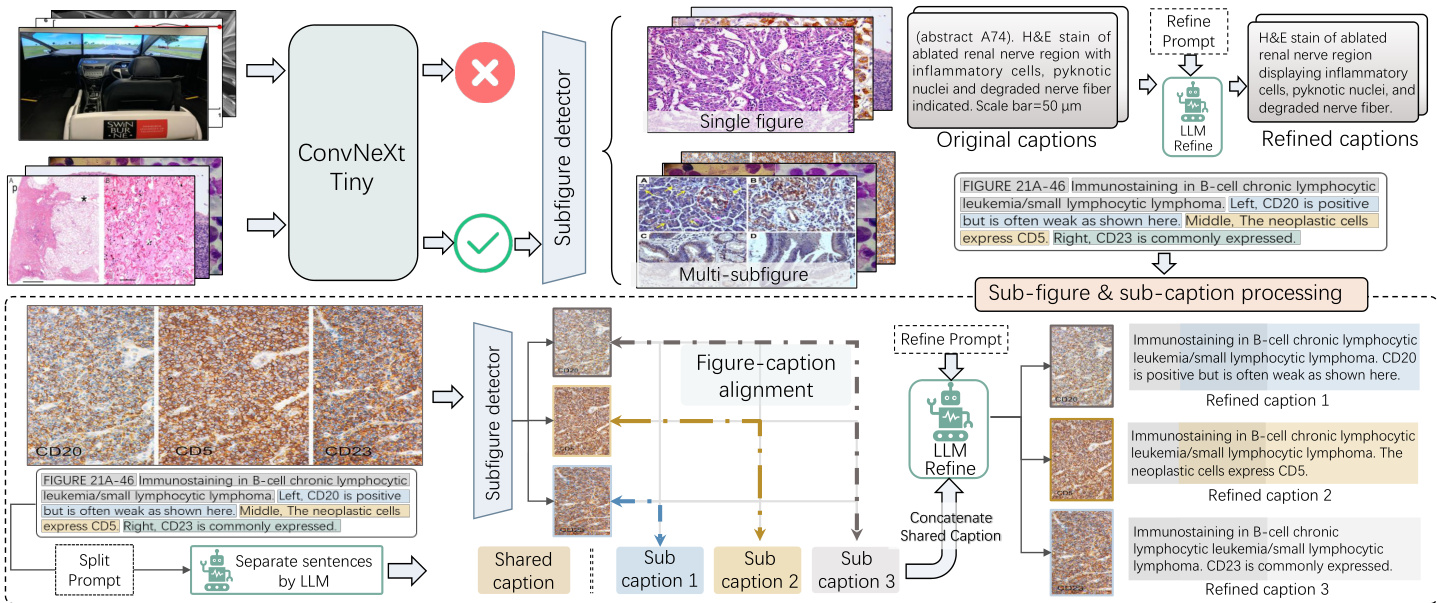
Figure 1: Illustration of data processing: pathology image selection, sub-figure & caption separation, and refinement.
图 1: 数据处理流程示意图:病理图像选择、子图与标题分离及精修。

Figure 2: Examples of pathology-specific model-invoking instruction-following samples.
图 2: 病理学专用模型调用指令遵循样本示例。
More specifically, data from PubMed are parsed from XML format papers into image-text pairs. For books, we first convert them from PDF to HTML and then parse the content into image-text pairs. Through these efforts, we collect 15M and 2K samples from these respective sources. Although the amount of data available on PubMed is substantial, it should be noted that the proportion of the data related to pathology is limited. Additionally, the clarity of these pathology images is comparatively inferior. Therefore, thorough filtering is required to ensure the quality and relevance of image-text pairs. As shown in Figure 1, our data cleansing process is executed methodically, following four carefully designed steps:
具体而言,我们从PubMed的XML格式论文中解析出图文对。对于书籍数据,首先将其从PDF转换为HTML格式,再从中提取图文对。通过上述方法,我们分别从这两个来源收集了1500万和2000条样本。虽然PubMed数据量庞大,但需注意其中病理学相关数据的占比有限,且这些病理图像的清晰度相对较低。因此需要通过严格筛选来确保图文对的质量与相关性。如图1所示,我们的数据清洗流程遵循四个精心设计的步骤有序执行:
Step 1: Pathology data selection. The dataset collected, especially from PubMed, encompasses a wide variety of image sources beyond the scope of pathology. To efficiently select pathology-related data, we manually annotate 20K samples, categorizing them as either pathological or nonpathological. Subsequently, we train a ConvNeXt (Liu et al. 2022a) model to identify pathological data within the remaining dataset, resulting in a pathology-specific dataset comprising 135K pathology-specific images.
步骤1:病理数据筛选。从PubMed等渠道收集的数据集包含大量病理学范畴之外的图像来源。为高效筛选病理相关数据,我们人工标注了2万份样本,将其分类为病理或非病理图像。随后训练ConvNeXt (Liu et al. 2022a)模型对剩余数据集进行病理数据识别,最终获得包含13.5万张病理图像的专用数据集。
Step 2: Sub-figure and sub-caption separation & alignment. In many instances, images consist of multiple subfigures, necessitating precise separation and alignment with their corresponding captions. As depicted in the lower half of Figure 1, we address the sub-figure separation by developing a YOLOv7 model (Wang, Boc hk ovsk iy, and Liao 2022) trained on 2K annotated bounding boxes. Regarding caption separation, conventional rule-based methods often fail to handle the separation of diverse and intricate captions. To overcome this limitation, we leverage the power of ChatGPT to automatically separate approximately 60K captions using carefully crafted prompts. Subsequently, we employ PLIP (Huang et al. 2023) to align sub-image with its corresponding sub-caption by assessing the similarity of visual content and captions. Moreover, we eliminate images with lower resolution, and remove the less relevant image-text pairs, further enhancing the overall quality of the dataset. Ultimately, we acquire 195K high-quality image-text pairs.
步骤2:子图与子标题的分离与对齐。在许多情况下,图像包含多个子图,需要精确分离并与对应标题对齐。如图1下半部分所示,我们通过训练基于2K标注边界框的YOLOv7模型(Wang, Bochkovskiy, and Liao 2022)解决子图分离问题。针对标题分离,传统基于规则的方法通常难以处理多样复杂的标题拆分。为此,我们利用ChatGPT的强大能力,通过精心设计的提示词自动分离约60K条标题。随后采用PLIP(Huang et al. 2023)通过评估视觉内容与标题的相似度实现子图与子标题的对齐。此外,我们移除低分辨率图像及关联性较弱的图文对,最终获得195K组高质量图文对。
Step 3: Caption refinement. As original captions include irrelevant information such as age and disease descriptions, and are not presented in a descriptive style. We design prompts to employ ChatGPT in refining the captions, making them more suitable for training.
步骤3:标题优化。由于原始标题包含年龄和疾病描述等无关信息,且未采用描述性风格呈现。我们设计提示词,利用ChatGPT优化标题,使其更适合训练。
Step 4: Instruction-following data generation. In this step, we select image-text pairs with captions exceeding 12 words. Using these pairs, we produce two types of instruction-following data: detailed description-based and conversation-based. The former is created by applying multiple well-designed instructions that inquire about detailed information, while the latter involves using ChatGPT to generate conversational Q&As based on the captions. Additionally, we design special model-invoking instructionfollowing samples covering a diverse range of scenarios, as depicted in Figure 2, enabling PathAsst with the capability to appropriately utilize pathology-specific sub-models.
步骤4: 指令跟随数据生成。本步骤筛选描述文本超过12词的图文配对数据,生成两类指令跟随数据:基于细节描述的指令数据和基于对话的指令数据。前者通过设计多组询问细节信息的指令模板生成,后者则利用ChatGPT根据描述文本生成对话式问答。此外,我们还设计了涵盖多样化场景的特殊模型调用指令样本 (如图2所示) ,使PathAsst具备合理调用病理学子模型的能力。
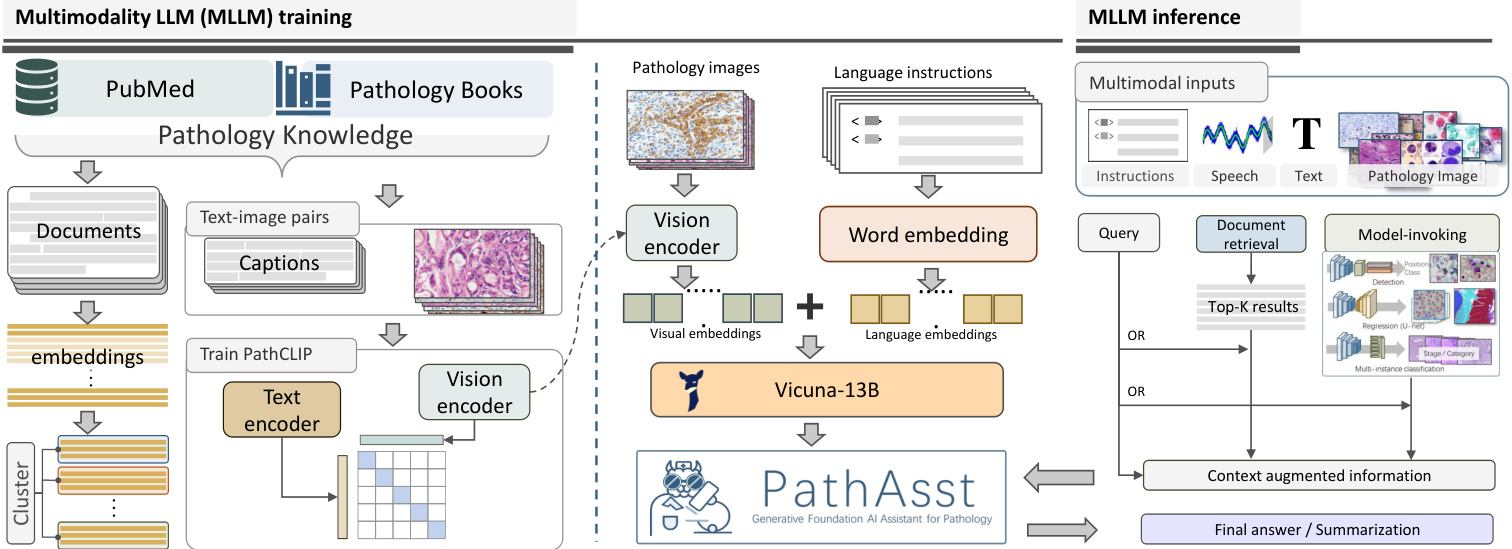
Figure 3: An illustration of the overall framework of PathAsst. The multimodal MLLM training encompasses the training processes of both PathCLIP and PathAsst, as well as the construction of a paper embedding database. The tool-augmented MLLM inference details the process of PathAsst utilizing various tools to enhance the quality of its generated outputs.
图 3: PathAsst整体框架示意图。多模态大语言模型训练包含PathCLIP和PathAsst的训练流程以及论文嵌入数据库的构建过程。工具增强的大语言模型推理详细描述了PathAsst利用各类工具提升生成输出质量的工作机制。
PathAsst Framework Construction
PathAsst框架构建
In this section, we present a comprehensive description of the construction process of PathAsst. This includes the introduction to the design of the model’s structure, training methodology, and the tools used for augmented model inference. A general overview can be found in Figure 3.
在本节中,我们将全面介绍PathAsst的构建过程,包括模型结构设计、训练方法以及用于增强模型推理的工具。整体概述可参见图3。
Model Design and Training
模型设计与训练
PathAsst is designed to integrate the strengths of both the advanced LLM and the CLIP (Radford et al. 2021) vision encoder to enable enhanced pathological analysis. For the visual component, we employ our custom-trained PathCLIP, complemented by a fully connected (FC) layer. Concerning the LLM component, we utilize Vicuna-13B (Chiang et al. 2023), a model widely recognized as the closest to ChatGPT in terms of performance. To elaborate, when an input image is provided, it is first encoded into visual tokens via the PathCLIP. Subsequently, the FC layer maps the image embedding space to the corresponding language embedding space. Finally, both visual and language embeddings are concatenated to the inputs of the MLLM. In the following, we introduce the detailed training process of PathCLIP and PathAsst.
PathAsst旨在整合先进大语言模型和CLIP (Radford et al. 2021) 视觉编码器的优势,以增强病理分析能力。视觉模块采用我们定制训练的PathCLIP,并辅以全连接 (FC) 层。大语言模型组件选用性能最接近ChatGPT的Vicuna-13B (Chiang et al. 2023)。具体而言,输入图像首先通过PathCLIP编码为视觉token,随后FC层将图像嵌入空间映射到对应的语言嵌入空间。最终,视觉与语言嵌入会被拼接为多模态大语言模型的输入。下文将详细介绍PathCLIP和PathAsst的训练流程。
Training of PathCLIP. As one of the core components of PathAsst, the capability of CLIP in interpreting pathological images largely dictates the performance ceiling of PathAsst. Therefore, we develop PathCLIP, a specialized variant of CLIP tailored for pathology. The training process involves fine-tuning a pre-trained OpenAI CLIP base model (Radford et al. 2021) using our PathCap dataset in a contrastive learning approach, following the training procedure from OpenCLIP repository (Ilharco et al. 2021). To be specific, for a batch of $N$ image-text pairs, PathCLIP is designed to maximize the cosine similarity between the embeddings of the pathology image and its corresponding text within each batch. Concurrently, it minimizes the cosine similarity amongst the remaining ${\dot{N}}^{2}-N$ non-pair samples. This strategy aligns the pathology vision and language space, thereby endowing PathCLIP with a more effective interpretation and analysis of pathology images.
PathCLIP的训练。作为PathAsst的核心组件之一,CLIP在病理图像解读方面的能力很大程度上决定了PathAsst的性能上限。为此,我们开发了PathCLIP——一个专为病理学定制的CLIP变体。训练过程采用对比学习方法,基于预训练的OpenAI CLIP基础模型(Radford等人,2021)和我们的PathCap数据集进行微调,遵循OpenCLIP代码库(Ilharco等人,2021)的训练流程。具体而言,对于包含$N$个图文对的批次,PathCLIP旨在最大化每个批次内病理图像嵌入与其对应文本嵌入之间的余弦相似度,同时最小化剩余${\dot{N}}^{2}-N$个非配对样本间的余弦相似度。该策略实现了病理视觉与语言空间的对齐,从而使PathCLIP能更有效地解读和分析病理图像。
Training of PathAsst. PathAsst is trained using the Path Instruct dataset through a two-phase training. In the first phase, both the vision encoder and the LLM are frozen, and we only train the FC layer that connects to the vision encoder. This initial phase aims to preliminarily align the vision encoder with the LLM. During this phase, we utilize the detailed description-based part of the Path Instruct. In the second phase, with an aspiration for PathAsst to generate higher-quality and more detailed responses, we extract all the data from books within the Path Instruct dataset, and include samples from PubMed with single images and captions exceeding the length of 50 tokens, resulting in a total training set of 35K samples. Only the PathCLIP is frozen during this phase’s training.
PathAsst的训练
PathAsst采用两阶段训练方式,基于Path Instruct数据集进行训练。第一阶段冻结视觉编码器和大语言模型,仅训练连接视觉编码器的全连接层(FC),旨在初步实现视觉编码器与大语言模型的对齐。此阶段使用Path Instruct中基于详细描述的数据部分。第二阶段,为提升PathAsst生成响应的高质量与细节丰富度,我们提取Path Instruct数据集中所有书籍数据,并加入来自PubMed的单图像样本(其标题长度超过50个token),最终构成35K样本的训练集。该阶段训练仅冻结PathCLIP模型。
Specifically, we standardize both forms of instructfollowing data formats, as shown in Table 1. First, we predefine a system message that sets the context for the LLM role. This is followed by a conversation between the user and the assistant, where the user provides instructions, and the assistant responds accordingly based on the instructions. To finetune our model, we utilize instruction-tuning via next-word prediction. Specifically, the model is trained to optimize the likelihood of generating an accurate response given the input image $\mathcal{T}$ and instruction $\mathbf{X}_ {i n s t r u c t}$ . The loss is calculated using the negative log-likelihood of the correct next token in the sequence, with the total loss summed across all time steps, which can be formulated as:
具体来说,我们对两种指令跟随数据格式进行了标准化处理,如表 1 所示。首先,我们预定义一个系统消息,用于设定大语言模型的角色背景。随后是用户与助手之间的对话,用户提供指令,助手根据指令做出相应回应。为了微调模型,我们通过下一词预测进行指令调优。具体而言,模型训练目标是优化在给定输入图像 $\mathcal{T}$ 和指令 $\mathbf{X}_ {instruct}$ 条件下生成准确响应的概率。损失函数采用序列中正确下一个 token 的负对数似然,所有时间步的总损失可表示为:
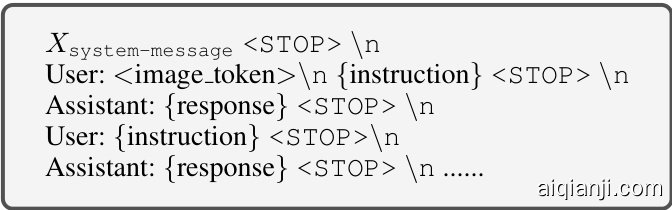

Table 1: Illustration of instruction-following data format, where {instruction} represents the user query, {response} denotes the corresponding answer. The $X_ {\tt s y s t e m-m e s s a g e}$ is set as: A dialogue between a professional pathology assistant and a human. The assistant provide informative, helpful, and detailed answers. The ${<}\mathrm{STOP>}$ is represented by ###, while stands for the tokens corresponding to the image tokens. During the model training, only {response} is considered when calculating loss.

表 1: 指令遵循数据格式示例,其中 {instruction} 表示用户查询,{response} 为对应回答。$X_ {\tt system-message}$ 设置为:专业病理助理与人类之间的对话,助理需提供信息丰富、有帮助且详尽的回答。${<}\mathrm{STOP>}$ 用 ### 表示, 代表图像对应的 token。模型训练时仅计算 {response} 部分的损失。
$$
\mathcal{L}(\pmb{\theta})=-\sum_ {t=1}^{T}\log p\left(x_ {t}\mid\mathcal{T},\mathbf{X}_ {i n s t r u c t},\mathbf{X}_ {\mathrm{a,<}t};\pmb{\theta}\right),
$$
$$
\mathcal{L}(\pmb{\theta})=-\sum_ {t=1}^{T}\log p\left(x_ {t}\mid\mathcal{T},\mathbf{X}_ {i n s t r u c t},\mathbf{X}_ {\mathrm{a,<}t};\pmb{\theta}\right),
$$
Where $\mathbf{X}_ {\mathrm{a},<t}$ refers to the prior tokens in the response sequence, $\theta$ denotes the trainable parameters of PathAsst. Specifically, during the first phase of training, $\theta$ corresponds to the parameters of the FC layer. In the subsequent phase, it represents both FC layer and LLM parameters. Meanwhile, $T$ signifies the length of the ground-truth response, and $p\left(x_ {t}\mid\bar{\mathcal{T}},\mathbf{X}_ {i n s t r u c t},\bar{\mathbf{X}}_ {\mathrm{a,<}t};\pmb{\theta}\right)$ represents the probability of generating the $t$ -th token in the response sequence.
其中 $\mathbf{X}_ {\mathrm{a},<t}$ 指代响应序列中的先前token,$\theta$ 表示PathAsst的可训练参数。具体而言,在训练的第一阶段,$\theta$ 对应全连接层(FC)的参数;在后续阶段,它同时代表全连接层和大语言模型(LLM)的参数。与此同时,$T$ 表示真实响应序列的长度,$p\left(x_ {t}\mid\bar{\mathcal{T}},\mathbf{X}_ {i n s t r u c t},\bar{\mathbf{X}}_ {\mathrm{a,<}t};\pmb{\theta}\right)$ 表示生成响应序列中第 $t$ 个token的概率。
Tool Augmented MLLM Inference
工具增强型多模态大语言模型推理
To augment PathAsst’s capabilities and offer more precise responses, we prepare two types of tools that PathAsst can employ during its inference phase. One leverages pathologyspecific computer vision (CV) sub-models, while the other focuses on paper retrieval. These tools not only enrich the context for PathAsst but also enable tasks beyond text generation, such as image generation and segmentation.
为增强PathAsst的能力并提供更精准的响应,我们为其推理阶段准备了两类工具:一类基于病理学专用计算机视觉(CV)子模型,另一类专注于论文检索。这些工具不仅能丰富PathAsst的上下文信息,还能实现文本生成之外的图像生成与分割等任务。
Pathology-specific CV Model Zoo. We integrate eight specialized pathological models into PathAsst for seamless invocation: (1) LBC (liquid-based cytology) classification model: This model is based on ConvNeXt-Tiny (Liu et al. 2022a), specifically designed for liquid-based cervical cytology image classification. Through the analysis of abnormal cell morph o logie s within the image, it effectively classifies the image into one of the six categories as defined by The Bethesda System (TBS). (2) LBC detection model: We utilize YOLOv7 (Wang, Boc hk ovsk iy, and Liao 2022) as the backbone for developing our detection model, which is employed to identify abnormal cells within image patches. This model is specifically designed to detect the five classes of non-normal cells as defined in TBS. (3) Hematological cell detection model: This model, developed based on YOLOv7, specializes in blood cell classification, which is crucial for diagnosing various hematological conditions. (4) LBC cell generation model: This model is developed based on Stable Diffusion (Rombach et al. 2022), which is capable of generating specific cells based on user input, such as ‘generate an image of a cell with nuclei enlarged 2-2.5 times’. (5) HER2 detection model, (6) PD-L1 detection model and (7) Ki67 detection model are developed using DPA-P2PNet (Shui et al. 2023) for immuno his to chemical cell detection and classification. (8) General segmentation model: Benefiting from the outstanding general segmentation quality of the Segment Anything Model (Kirillov et al. 2023), we directly employ it as our pathology image segmentation model.
病理专用CV模型库。我们整合了八种专业病理模型到PathAsst中实现无缝调用:(1) LBC(液基细胞学)分类模型:该模型基于ConvNeXt-Tiny (Liu et al. 2022a),专为液基宫颈细胞学图像分类设计。通过分析图像中的异常细胞形态学特征,可有效将图像分类为TBS(The Bethesda System)定义的六种类别之一。(2) LBC检测模型:采用YOLOv7 (Wang, Bochkovskiy, and Liao 2022)作为主干网络开发的检测模型,用于识别图像区块中的异常细胞,专门检测TBS定义的五类非正常细胞。(3) 血细胞检测模型:基于YOLOv7开发,专攻血细胞分类,对诊断各类血液病症至关重要。(4) LBC细胞生成模型:基于Stable Diffusion (Rombach et al. 2022)开发,能根据用户输入生成特定细胞,例如"生成细胞核增大2-2.5倍的细胞图像"。(5) HER2检测模型、(6) PD-L1检测模型和(7) Ki67检测模型均采用DPA-P2PNet (Shui et al. 2023)开发,用于免疫组织化学细胞检测与分类。(8) 通用分割模型:得益于Segment Anything Model (Kirillov et al. 2023)卓越的通用分割质量,我们直接将其作为病理图像分割模型使用。
Table 2: Comparative evaluation of zero-shot image classification performance across different CLIP models.
| Model | CRC | WSSS4LUAD | LC-lung | LC-colon |
| OpenAI CLIP | 22.2 | 61.6 | 31.5 | 75.7 |
| PLIP | 53.1 | 69.5 | 86.0 | 87.0 |
| PathCLIP | 54.2 | 81.1 | 88.7 | 94.3 |
表 2: 不同CLIP模型在零样本图像分类任务中的性能对比评估
| 模型 | CRC | WSSS4LUAD | LC-lung | LC-colon |
|---|---|---|---|---|
| OpenAI CLIP | 22.2 | 61.6 | 31.5 | 75.7 |
| PLIP | 53.1 | 69.5 | 86.0 | 87.0 |
| PathCLIP | 54.2 | 81.1 | 88.7 | 94.3 |
Once PathAsst invokes a particular specialized model, it processes both the user’s query and the output of the invoked model to formulate a conclusive response, resulting in a more precise and effective interaction with the user.
当PathAsst调用某个特定专业模型时,它会同时处理用户的查询和被调用模型的输出,以形成最终结论性响应,从而实现与用户更精准高效的交互。
Enhancing Responses through Paper Retrieval. In the realm of pathology, even the highly recognized GPT-4 struggles with specific queries that necessitate deep domain knowledge, especially apparent when addressing questions involving the most recent research. Taking inspiration from Langchain’s approach (Chase 2022) for building local knowledge databases, we gather $5.3\mathrm{M}$ article abstracts from PubMed. We utilize PubMedBERT (Gu et al. 2021) for abstract embedding extraction and Faiss (Johnson, Douze, and Jégou 2019) for the efficient storage of these embeddings. To expedite inference efficiency, a preliminary abstract clustering is conducted. Upon user query, our system allows the extraction of relevant information from this paper database, serving as context information to amplify the precision of LLM’s responses.
通过论文检索增强回答能力。在病理学领域,即使是备受推崇的GPT-4也难以应对需要深厚领域知识的特定查询,尤其在涉及最新研究的问题时表现明显。受Langchain构建本地知识库方法(Chase 2022)的启发,我们从PubMed收集了530万篇论文摘要,使用PubMedBERT(Gu等人2021)进行摘要嵌入提取,并采用Faiss(Johnson、Douze和Jégou 2019)高效存储这些嵌入向量。为提升推理效率,系统会预先对摘要进行聚类处理。当用户发起查询时,我们的系统能够从该论文数据库中提取相关信息,作为上下文信息来提升大语言模型回答的精准度。
Experiments
实验
Evaluation Datasets Construction. We construct and gather a series of test datasets to evaluate the performance of the proposed PathCLIP and PathAsst.
评估数据集构建。我们构建并收集了一系列测试数据集,用于评估所提出的PathCLIP和PathAsst的性能。
For the evaluation of zero-shot classification of PathCLIP, we collect: (1) CRC100K dataset (Kather, Halama, and Marx 2018): This is a collection of 100K image patches derived from H&E stained his to logical images of both colorectal cancer and normal tissue, categorized into nine tissue classes, including Adipose, Background, Debris, Lymphocytes, Mucus, Smooth Muscle, Normal Colon Mucosa, Cancer-Associated Stroma, and Colorectal A de no carcinoma Epithelium. (2) WSSS4LUAD (Han et al. 2022): This dataset comprises patch-level annotations from 87 whole slide images. In this case, we focus on the tumor and normal classes, which yields a total of 6,579 tumor and 1,832 normal images. In order to assess the model on these datasets, labels are transformed into complete sentences. For instance, the label ‘tumor’ is rephrased as ‘A H&E image of a tumor.’ (3) LC25000 (Borkowski et al. 2019). This dataset comprises tissue samples from lung and colon adenocarcinomas, divided into two distinct subsets: the LC-lung and the LC-colon. The LC-lung encompasses 15,000 images and includes classifications of lung a de no carcinomas, lung squamous cell carcinomas, and benign lung tissues. On the other hand, the LC-colon subset, containing 10,000 images, is categorized into colon a de no carcinomas and benign colonic tissues. The F1 score is used as the metric for evaluation.
为评估PathCLIP的零样本分类能力,我们收集了以下数据集:(1) CRC100K数据集(Kather, Halama, and Marx 2018):该数据集包含10万张从结直肠癌及正常组织的H&E染色组织学图像中提取的切片,分为九类组织类型,包括脂肪组织、背景、碎片、淋巴细胞、粘液、平滑肌、正常结肠黏膜、癌相关间质和结直肠腺癌上皮。(2) WSSS4LUAD(Han et al. 2022):该数据集包含87张全切片图像的区块级标注,我们仅关注肿瘤与正常组织两类,共获得6,579张肿瘤图像和1,832张正常图像。为适配模型评估,标签被转换为完整句子,例如将"tumor"改写为"A H&E image of a tumor"。(3) LC25000(Borkowski et al. 2019):该数据集包含肺腺癌和结肠腺癌组织样本,分为LC-lung和LC-colon两个子集。LC-lung子集包含15,000张图像,涵盖肺腺癌、肺鳞状细胞癌和良性肺组织;LC-colon子集包含10,000张图像,分为结肠腺癌和良性结肠组织。评估采用F1分数作为指标。

Figure 4: Comparative assessment of image retrieval performance between CLIP models across collected datasets.
图 4: 在不同数据集上 CLIP 模型的图像检索性能对比评估

Figure 5: Example of PathAsst calls generation model.
图 5: PathAsst 调用生成模型的示例。
For the cross-modal retrieval validation of PathCLIP, we employ the test set from the PubMed section of our collected data, along with data from books. Note that the data from books are not included during the training phase of the PathCLIP, hence providing an evaluation in the context of unseen domains. Given that the length of some captions in these datasets is relatively long and may exceed the token length limitation of CLIP, we opt for samples with captions that are fewer than 77 tokens. The @kmetric is used to assess the performance of image retrieval, which measures whether the correct image is presented among the T opk retrieved images.
为验证PathCLIP的跨模态检索能力,我们采用收集数据中PubMed部分的测试集及图书数据。需注意图书数据未参与PathCLIP训练阶段,从而实现对未见领域的评估。鉴于部分数据集的描述文本较长可能超出CLIP的token长度限制(77个),我们筛选了描述文本少于77个token的样本。使用R@k指标评估图像检索性能,该指标衡量正确图像是否出现在Topk检索结果中。
For the validation of PathAsst, we employ the PathVQA dataset (He et al. 2020), which comprises 32,799 questions derived from 4,998 pathology images. The type of questions includes open-ended questions typically beginning with what, where, and when, as well as close-ended questions requiring yes/no responses. We measure model performance of close-ended questions using accuracy, and evaluate open-ended questions with F1-score.
为验证PathAsst,我们采用PathVQA数据集 (He et al. 2020),该数据集包含源自4,998张病理学图像的32,799个问题。问题类型包括通常以what、where和when开头的开放式问题,以及需要yes/no回答的封闭式问题。我们使用准确率衡量封闭式问题的模型性能,并以F1分数评估开放式问题。
Table 3: Comparison of various methods on PathVQA.
| Method | PathVQA | |
| Closed | Open | |
| M2I2 (Liet al.2022) | 88.0 | 36.3 |
| CLIP-ViTw/GPT2(vanSonsbeeket al.2023) | 87.0 | 40.0 |
| MMQ (Doetal.2021) | 84.0 | 13.4 |
| LLaVA (Liuet al.2023a) | 81.0 | 19.2 |
| BLIP-2Flan-T5XXL (Liet al.2023) | 80.1 | 34.1 |
| PathAsst (w/ CLIP) | 89.7 | 37.6 |
| PathAsst(w/PathCLIP) | 90.9 | 38.4 |
表 3: PathVQA上各种方法的比较
| 方法 | PathVQA |
|---|---|
| 封闭式 | |
| M2I2 (Li et al. 2022) | 88.0 |
| CLIP-ViTw/GPT2 (van Sonsbeek et al. 2023) | 87.0 |
| MMQ (Do et al. 2021) | 84.0 |
| LLaVA (Liu et al. 2023a) | 81.0 |
| BLIP-2 Flan-T5 XXL (Li et al. 2023) | 80.1 |
| PathAsst (w/ CLIP) | 89.7 |
| PathAsst (w/ PathCLIP) | 90.9 |
Statistical Results. As shown in Table 2 and Figure 4. Our analysis demonstrates that our PathCLIP significantly surpasses the baseline OpenAI CLIP model, consistently outperforming the state-of-the-art (SOTA) pathology model, PLIP, in tasks such as cross-modal image retrieval and zeroshot image classification. To be specific, PathCLIP achieves a remarkable improvement in the $R@\mathrm{10}$ retrieval on the PubMed dataset, with a 10.71-fold and 11.07-fold increase compared to the OpenAI CLIP and PLIP models, respectively. In the context of unseen domain data, the retrieval $R@\mathrm{10}$ on the books is 5.78 times and 2.38 times that of OpenAI CLIP and PLIP, respectively. Considering the zeroshot classification tasks, PathCLIP achieves a substantial improvement in F1-score compared to CLIP, with notable gains of $32%$ , $19.5%$ , $57.2%$ , and $18.6%$ on the CRC100K, WSSS4LUAD, LC-lung, and LC-colon datasets, respec- tively. Furthermore, even when compared to the previous SOTA PLIP model, PathCLIP shows an increase of $1.1%$ , $11.6%$ , $2.7%$ , and $7.3%$ on these datasets, respectively. For the evaluation on PathVQA, PathAsst significantly outperforms the prior MLLM model in both closed-form and openended question types. Specifically, it surpasses LLaVA by $8.7%$ and $18.4%$ in these two question types, respectively. This underscores the importance of training with PathInstruct data. Further enhancements of $1.2%$ and $0.8%$ are noted after substituting CLIP with PathCLIP, indicating that
统计结果。如表 2 和图 4 所示,我们的分析表明,PathCLIP 显著超越了基线 OpenAI CLIP 模型,在跨模态图像检索和零样本图像分类等任务中持续优于当前最优 (SOTA) 病理学模型 PLIP。具体而言,PathCLIP 在 PubMed 数据集上的 $R@\mathrm{10}$ 检索实现了显著提升,相比 OpenAI CLIP 和 PLIP 模型分别提高了 10.71 倍和 11.07 倍。在未见领域数据方面,书籍检索的 $R@\mathrm{10}$ 分别是 OpenAI CLIP 和 PLIP 的 5.78 倍和 2.38 倍。在零样本分类任务中,PathCLIP 的 F1 分数相比 CLIP 有显著提升,在 CRC100K、WSSS4LUAD、LC-lung 和 LC-colon 数据集上分别提高了 $32%$、$19.5%$、$57.2%$ 和 $18.6%$。此外,即使与之前的 SOTA PLIP 模型相比,PathCLIP 在这些数据集上仍分别提升了 $1.1%$、$11.6%$、$2.7%$ 和 $7.3%$。在 PathVQA 评估中,PathAsst 在封闭式和开放式问题类型上均显著优于先前的 MLLM 模型,具体而言,在这两类问题中分别超越了 LLaVA $8.7%$ 和 $18.4%$,这凸显了使用 PathInstruct 数据训练的重要性。在用 PathCLIP 替换 CLIP 后,性能进一步提升了 $1.2%$ 和 $0.8%$,表明


Provide a comprehensive analysis of the findings shown in the image.
对图中所示结果进行全面分析。
PathAsst
PathAsst
Llava
Llava
Mini-GPT4
Mini-GPT4

Figure 6: An example of PathAsst invokes the PD-L1 detection model for assistance. Figure 7: An example of PathAsst, LLaVA, and MiniGPT-4’s capability in interpreting pathology images.
图 6: PathAsst 调用 PD-L1 检测模型进行辅助的示例。图 7: PathAsst、LLaVA 和 MiniGPT-4 在病理图像解读能力上的对比示例。
Based on these observations. I would recommend further testing to confirm the presence of abnormal cells. An abnormal cervical Pap test result may indicate a hich-risk for cervical cancer. It's important to follow up with your healthcare provider for further evaluation and treatment.
基于这些观察结果,建议进一步检测以确认异常细胞的存在。异常的宫颈巴氏涂片检查结果可能提示宫颈癌高风险。请务必联系您的医疗服务提供者进行后续评估和治疗。
The image is a close-up view of a blue substance, possibly ink or a dye, dispersed in water. The blue particles are evenly distributed throughout the water, creating a visually striking scene. The blue substance appears to be in the form of small spheres or shapes, which are spread across the entire image. The contrast between the blue particles and the water makes the scene quite captivating, as the blue elements stand out prominently.
图片展示了一种蓝色物质(可能是墨水或染料)在水中扩散的特写画面。蓝色颗粒均匀分布在水中,形成极具视觉冲击力的场景。这些蓝色物质呈现为小球状或各种形状,遍布整个画面。蓝色颗粒与水体的鲜明对比使整个场景格外引人注目,蓝色元素在画面中尤为突出。
The image shows a group of cells, which are microscopic in size, arranged in a pattern. The cells are blue and have a round shape. Some of the cells have small black dots on them, which could be the nuclei. The cells appear to be arranged in a grid-like pattern, with some cells connected to others. The cells are suspended in a clear liquid, which could be a solution. The image is likely a microscopic view of a sample of cells.
图中展示了一组微观尺寸的细胞,呈规则排列。细胞呈蓝色圆形,部分细胞表面带有小黑点(可能是细胞核)。这些细胞以网格状排布,部分细胞相互连接。细胞悬浮在透明液体中,可能是某种溶液。该图像很可能是细胞样本的显微视图。
the incorporation of PathCLIP enhances PathAsst’s understanding of pathology images. Compared with the performance of the previous SOTA model, which directly extracts the statistical number from their reports, PathAsst achieves considerable improvements in closed-ended questions, although it slightly under performs the SOTA model in openended questions.
PathCLIP的整合增强了PathAsst对病理图像的理解能力。相比之前直接从报告中提取统计数据的SOTA模型,PathAsst在封闭式问题上取得了显著提升,尽管在开放式问题上表现略逊于SOTA模型。
Demonstration Showcase of PathAsst. Here, we showcase several examples of PathAsst’s robust capabilities in handling complex pathology tasks. As shown in Figure 5, PathAsst is capable of recognizing the user’s need to generate an LBC cell that belongs to the LSIL category with irregular excavated cytoplasm. It accomplishes this by invoking the LBC cell generation model. This advanced functionality empowers users to create a diverse range of LBC cells that are precisely tailored to their specific needs.
PathAsst 功能演示展示。在此,我们展示几个 PathAsst 在处理复杂病理学任务方面的强大能力示例。如图 5 所示,PathAsst 能够识别用户生成属于 LSIL 类别、具有不规则凹陷细胞质的 LBC 细胞的需求,并通过调用 LBC 细胞生成模型实现这一功能。这一先进功能使用户能够创建多种多样的 LBC 细胞,精准满足其特定需求。
Figure 6 illustrates another example of PathAsst employing a model invocation, where the user requires to count the positive cells in the image, which can be challenging through direct multimodal generation. Therefore, PathAsst chooses to invoke the PD-L1 cell detection model. It automatically marks the predicted points on the cells in the image and provides the statistical results for further analysis with LLM. In this case, LLM generates a markdown-formatted table to display the results along with the corresponding analysis.
图 6: 展示了PathAsst采用模型调用的另一个示例。用户需要统计图像中的阳性细胞数量,这通过直接多模态生成可能具有挑战性。因此,PathAsst选择调用PD-L1细胞检测模型,自动在图像中的细胞上标记预测点,并提供统计结果供大语言模型进一步分析。本例中,大语言模型生成Markdown格式的表格来展示结果及相应分析。
Furthermore, Figure 7 demonstrates PathAsst’s ability to interpret pathology images independently. In comparison to LLaVA and MiniGPT-4, PathAsst places greater emphasis on cell morphology and features, such as enlarged nucleus and irregular nuclear membrane. In contrast, LLaVA fails to recognize the image as pathological, while MiniGPT-4 generates simplistic descriptions such as ‘cells are blue and have a round shape’ and ‘cells are suspended in a clear liquid.’
此外,图7展示了PathAsst独立解读病理图像的能力。与LLaVA和MiniGPT-4相比,PathAsst更注重细胞形态和特征,例如增大的细胞核和不规则的核膜。相比之下,LLaVA未能识别该图像为病理图像,而MiniGPT-4仅生成简单描述,如"细胞呈蓝色且形状圆润"和"细胞悬浮在透明液体中"。
Conclusion
结论
In this study, we construct PathCap and Path Instruct datasets, comprising 207K pathology image-text pairs and 180K instruction-following samples, by systematically collecting and processing pathology data from various sources. Leveraging these high-quality datasets, we propose PathCLIP and PathAsst. PathCLIP exhibits powerful capabilities in pathology cross-modal retrieval and zero-shot classification. PathAsst, an instruction-tuned foundation model, is a synergy of the powerful vision encoder PathCLIP and the Vicuna-13b LLM, equipped with an established toolkit that includes eight pathology-specific models and a 5.3 millionsized paper retrieval system. PathAsst not only showcases impressive capabilities in pathology multimodal dialogue and interpreting pathology images, but also the ability to handle more complex pathology tasks by the invocation of these established pathology tools. We hope that the construction of model frameworks and datasets can offer insights and aid in the advancement of pathology foundational models.
在本研究中,我们通过系统收集和处理来自不同来源的病理学数据,构建了PathCap和Path Instruct数据集,包含207K病理图像-文本对和180K指令遵循样本。利用这些高质量数据集,我们提出了PathCLIP和PathAsst。PathCLIP在病理学跨模态检索和零样本分类方面展现出强大能力。PathAsst是一个经过指令调优的基础模型,结合了强大的视觉编码器PathCLIP和Vicuna-13b大语言模型,并配备了一个成熟的工具包,其中包括八个病理学专用模型和一个530万规模的论文检索系统。PathAsst不仅在病理学多模态对话和解释病理图像方面展现出令人印象深刻的能力,还能通过调用这些成熟的病理学工具来处理更复杂的病理学任务。我们希望模型框架和数据集的构建能为病理学基础模型的进步提供见解和帮助。
Acknowledgements
致谢
This study was partially supported by the National Natural Science Foundation of China (Grant No.92270108), Zhejiang Provincial Natural Science Foundation of China (Grant No.XHD23F0201), and the Research Center for Industries of the Future (RCIF) at Westlake University.
本研究部分由国家自然科学基金(资助号:92270108)、浙江省自然科学基金(资助号:XHD23F0201)以及西湖大学未来产业研究中心支持。
References Alayrac, J.-B.; Donahue, J.; Luc, P.; Miech, A.; Barr, I.; Has- son, Y.; Lenc, K.; Mensch, A.; Millican, K.; Reynolds, M.; et al. 2022. Flamingo: a Visual Language Model for FewShot Learning. In Advances in Neural Information Processing Systems. Borkowski, A. A.; Bui, M. M.; Thomas; et al. 2019. Lung and colon cancer his to pathological image dataset (lc25000). arXiv preprint arXiv:1912.12142. Brown, T.; Mann, B.; Ryder, N.; Subbiah, M.; Kaplan, J. D.; Dhariwal, P.; Neel a kant an, A.; Shyam, P.; Sastry, G.; Askell, A.; et al. 2020. Language models are few-shot learners. Advances in neural information processing systems, 33: 1877– 1901. Changpinyo, S.; Sharma, P.; Ding, N.; and Soricut, R. 2021. Conceptual $12\mathrm{m}$ : Pushing web-scale image-text pre-training to recognize long-tail visual concepts. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 3558–3568. Chase, H. 2022. LangChain. Chiang, W.-L.; Li, Z.; Lin, Z.; Sheng, Y.; Wu, Z.; Zhang, H.; Zheng, L.; Zhuang, S.; Zhuang, Y.; Gonzalez, J. E.; Stoica, I.; and Xing, E. P. 2023. Vicuna: An Open-Source Chatbot Impressing GPT-4 with $90%* $ ChatGPT Quality. Chowdhery, A.; Narang, S.; Devlin, J.; Bosma, M.; Mishra, G.; Roberts, A.; Barham, P.; Chung, H. W.; Sutton, C.; Gehrmann, S.; et al. 2022. Palm: Scaling language modeling with pathways. arXiv preprint arXiv:2204.02311. Chung, H. W.; Hou, L.; Longpre, S.; Zoph, B.; Tay, Y.; Fe-dus, W.; Li, E.; Wang, X.; Dehghani, M.; Brahma, S.; et al.2022. Scaling instruction-finetuned language models. arXiv preprint arXiv:2210.11416. Devlin, J.; Chang, M.-W.; Lee, K.; and Toutanova, K. 2018. Bert: Pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805. Do, T.; Nguyen, B. X.; Tjiputra, E.; Tran, M.; Tran, Q. D.; and Nguyen, A. 2021. Multiple meta-model quantifying for medical visual question answering. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part V 24, 64–74. Springer. Driess, D.; Xia, F.; Sajjadi, M. S.; Lynch, C.; Chowdhery, A.; Ichter, B.; Wahid, A.; Tompson, J.; Vuong, Q.; Yu, T.; et al. 2023. Palm-e: An embodied multimodal language model. arXiv preprint arXiv:2303.03378. Gao, P.; Han, J.; Zhang, R.; Lin, Z.; Geng, S.; Zhou, A.; Zhang, W.; Lu, P.; He, C.; Yue, X.; et al. 2023. LLaMAAdapter V2: Parameter-Efficient Visual Instruction Model. arXiv preprint arXiv:2304.15010.
参考文献
Alayrac, J.-B.; Donahue, J.; Luc, P.; Miech, A.; Barr, I.; Hasson, Y.; Lenc, K.; Mensch, A.; Millican, K.; Reynolds, M.; 等. 2022. Flamingo: 一种用于少样本学习的视觉语言模型. 见《神经信息处理系统进展》.
Borkowski, A. A.; Bui, M. M.; Thomas; 等. 2019. 肺和结肠癌组织病理学图像数据集 (lc25000). arXiv预印本 arXiv:1912.12142.
Brown, T.; Mann, B.; Ryder, N.; Subbiah, M.; Kaplan, J. D.; Dhariwal, P.; Neelakantan, A.; Shyam, P.; Sastry, G.; Askell, A.; 等. 2020. 语言模型是少样本学习者. 《神经信息处理系统进展》, 33: 1877–1901.
Changpinyo, S.; Sharma, P.; Ding, N.; Soricut, R. 2021. Conceptual $12\mathrm{m}$: 将网络规模图文预训练推向长尾视觉概念识别. 见《IEEE/CVF计算机视觉与模式识别会议论文集》, 3558–3568.
Chase, H. 2022. LangChain.
Chiang, W.-L.; Li, Z.; Lin, Z.; Sheng, Y.; Wu, Z.; Zhang, H.; Zheng, L.; Zhuang, S.; Zhuang, Y.; Gonzalez, J. E.; Stoica, I.; Xing, E. P. 2023. Vicuna: 一款以$90%* $ ChatGPT质量惊艳GPT-4的开源聊天机器人.
Chowdhery, A.; Narang, S.; Devlin, J.; Bosma, M.; Mishra, G.; Roberts, A.; Barham, P.; Chung, H. W.; Sutton, C.; Gehrmann, S.; 等. 2022. Palm: 基于Pathways扩展的语言建模. arXiv预印本 arXiv:2204.02311.
Chung, H. W.; Hou, L.; Longpre, S.; Zoph, B.; Tay, Y.; Fedus, W.; Li, E.; Wang, X.; Dehghani, M.; Brahma, S.; 等. 2022. 扩展指令微调语言模型. arXiv预印本 arXiv:2210.11416.
Devlin, J.; Chang, M.-W.; Lee, K.; Toutanova, K. 2018. Bert: 面向语言理解的深度双向Transformer预训练. arXiv预印本 arXiv:1810.04805.
Do, T.; Nguyen, B. X.; Tjiputra, E.; Tran, M.; Tran, Q. D.; Nguyen, A. 2021. 医学视觉问答的多元模型量化. 见《医学图像计算与计算机辅助介入–MICCAI 2021: 第24届国际会议论文集, 法国斯特拉斯堡》, 64–74. Springer.
Driess, D.; Xia, F.; Sajjadi, M. S.; Lynch, C.; Chowdhery, A.; Ichter, B.; Wahid, A.; Tompson, J.; Vuong, Q.; Yu, T.; 等. 2023. Palm-e: 具身多模态语言模型. arXiv预印本 arXiv:2303.03378.
Gao, P.; Han, J.; Zhang, R.; Lin, Z.; Geng, S.; Zhou, A.; Zhang, W.; Lu, P.; He, C.; Yue, X.; 等. 2023. LLaMAAdapter V2: 参数高效的视觉指令模型. arXiv预印本 arXiv:2304.15010.
