A guide to artificial intelligence for cancer researchers
癌症研究人员人工智能指南
Abstract
摘要
| Sections |
| Introduction |
| Understandingdeeplearning |
| Al for biomedical image analysis |
| Alforlanguage |
| Emerging usesofAl |
| Conclusion |
| 章节 |
|---|
| 引言 |
| 理解深度学习 |
| 用于生物医学图像分析的AI |
| 用于语言的AI |
| AI的新兴应用 |
| 结论 |
Artificial intelligence (Al) has been com mod it i zed. It has evolved from a specialty resource to a readily accessible tool for cancer researchers. Al-based tools can boost research productivity in daily workflows, but can also extract hidden information from existing data, thereby enabling new scientific discoveries. Building a basic literacy in these tools is useful for every cancer researcher. Researchers with a traditional biological science focus can use Al-based tools through off-the-shelf software, whereas those who are more computationally inclined can develop their own Al-based software pipelines. In this article, we provide a practical guide for non-computational cancer researchers to understand how Al-based tools can benefit them. We convey general principles of Al for applications in image analysis, natural language processing and drug discovery. In addition, we give examples of how non-computational researchers can get started on the journey to productively use Al in their own work.
人工智能 (AI) 已被商品化。它已从专业资源演变为癌症研究人员随手可得的工具。基于AI的工具既能提升日常研究工作流程的效率,也能从现有数据中提取隐藏信息,从而推动新的科学发现。掌握这些工具的基础应用能力对每位癌症研究者都大有裨益。专注于传统生物科学的研究人员可通过现成软件使用基于AI的工具,而更倾向于计算方向的研究者则可自行开发基于AI的软件流程。本文为非计算背景的癌症研究人员提供实用指南,帮助他们理解基于AI的工具如何惠及自身研究。我们将阐述AI在图像分析、自然语言处理和药物发现领域应用的通用原则,并举例说明非计算背景研究人员如何在本职工作中高效运用AI技术。
Review article
综述文章
Introduction
引言
Artificial intelligence (Al) is a set of computational techniques that aim to enable machines to perform tasks that are usually reserved for humans.For decades,Alhas been mostly a theoretical construct with few implications for the real world. However, in the past 15 years, Al has reached human-level performance in a broad range of domains, including biomedical research (Fig.1a and Box1).Alhas matured into a multitude of products with real-world implications. Technically, many of the present-day applications of Al heavily rely on artificial neural networks (ANNs), computational models inspired by the biological neural networks in the brain. An ANN is composed of linked units called'neurons', which are organized into layers. ANNs typically have multiple layers, each containing numerous neurons: an input layer, which receives the input data;an output layer, which produces the final output; and hidden layers, which perform most of the computation. When an ANN contains a substantial number of these hidden layers, it is referred to as a deep ANN.Theterm'deep'in this context relates to the depth of the network, which is a measure of the number of layers. Deep neural networks are capable oflearning complex patterns due to their extended architecture, which allows for more levels of abstraction and representation of data. Deep neural networks are trained using large datasets and adjusting the parameters of the network (such as the weights of connections between neurons) based on the output error. This process is known as deep learning. During deep learning, the network learns to perform tasks from examples. Today, in cancer research,the concept of Aloverlaps with deep learning to a high degree, such that these terms are often used interchangeably.
人工智能 (AI) 是一套旨在让机器执行通常由人类完成的任务的计算技术。几十年来,AI 主要是一种理论构建,对现实世界影响甚微。然而在过去 15 年间,AI 已在包括生物医学研究在内的广泛领域达到人类水平性能 (图 1a 和框 1)。AI 已发展成众多具有现实意义的产品。从技术角度而言,当今许多 AI 应用严重依赖人工神经网络 (ANN)——这种计算模型受大脑中生物神经网络启发。ANN 由称为"神经元"的互联单元组成,这些单元分层排列。ANN 通常具有多个层:接收输入数据的输入层;产生最终输出的输出层;以及承担主要计算任务的隐藏层。当 ANN 包含大量隐藏层时,即被称为深度 ANN。此处的"深度"指网络深度,即层数的度量。深度神经网络凭借其扩展架构能够学习复杂模式,这种架构支持更多层级的抽象和数据表示。深度神经网络使用大型数据集进行训练,并根据输出误差调整网络参数(如神经元连接权重)。这一过程被称为深度学习。在深度学习过程中,网络通过示例学习执行任务。当今在癌症研究中,AI 的概念与深度学习高度重合,因此这些术语经常互换使用。
Al methods are becoming increasingly common in cancer research. Hence, we postulate that any cancer researcher nowadays needs to acquire a certain level of Al literacy. Today, it is important to be able to understand, interpret and critically evaluate the Al output. In addition, some cancer researchers will find it beneficial to acquire a deeper understanding of Al and develop their own Al-based software tools. Today, Al has been com mod it i zed, meaning it is no longer a specialized resource but a widely accessible tool that cancer researchers can readily utilize (Fig.la). In addition, some biomedical research software features Al systems that can be trained for specific tasks,such as image analysis tools with a graphical user interface that allow users to train custom Al models(Fig.1b and Supplementary Table 1).Although a complete and detailed understanding of the exact mechanisms of Al systems remains a challenging and evolving area of research, it is important to recognize the limitations and potential biases inherent in these systems based on a general understanding of how Al systems are trained and how they make their predictions.
人工智能方法在癌症研究中正变得越来越普遍。因此,我们主张当今任何癌症研究人员都需要掌握一定程度的人工智能素养。如今,能够理解、解释和批判性评估人工智能输出已变得尤为重要。此外,部分癌症研究人员会发现深入理解人工智能并开发自己的人工智能软件工具将大有裨益。当前,人工智能已实现商品化 (commoditized) ,这意味着它不再是专业资源,而是癌症研究人员可随时使用的普及工具 (图1a) 。同时,部分生物医学研究软件配备了可针对特定任务训练的人工智能系统,例如具有图形用户界面的图像分析工具,允许用户训练定制化人工智能模型 (图1b和补充表1) 。尽管要完全详细理解人工智能系统的确切机制仍是一个不断发展的挑战性研究领域,但基于对人工智能系统训练方式及其预测机制的基本理解,认识这些系统固有的局限性和潜在偏差至关重要。
This Review aims to serve as a comprehensive guide for cancer researchers who want to deepen their understanding of the applications of Al in cancer research. Our experience in engaging with professionals across disciplines such as biology, medicine and biochemistry has highlighted agrowing need forclarity and guidance on Alconcepts and tools that are pertinent to cancer research. This Review focuses on the key concepts and tools related to Al in cancer research, including applications in image analysis, natural language processing (NLP) and drug discovery. It is intended as a practical guide and will not cover the mathematical foundations or highly technical aspects of Al algorithms in depth.
本综述旨在为希望深入了解人工智能 (AI) 在癌症研究中应用的科研人员提供全面指南。我们在与生物学、医学和生物化学等跨学科专业人士的合作中发现,研究人员对癌症研究相关的AI概念与工具存在日益增长的明晰化指导需求。本文重点关注癌症研究中AI相关的核心概念与工具,包括其在图像分析、自然语言处理 (NLP) 和药物发现领域的应用。本指南侧重实践性,不会深入探讨AI算法的数学基础或高技术性内容。
Understanding deep learning Types of deep learning
理解深度学习 深度学习的类型
Deep learning algorithms can be split into supervised', unsupervised? and reinforcement 34 learning. Supervised learning is the most common type of deep learning. In supervised learning, a model is trained on a labelled dataset, which means that each training example (forexample,a photo) ispaired with an output label or ground truth label (for example,'cat'or'dog).A model makes predictions or decisions based on the input data, and it learns patterns for the correct predictions from the ground-truth labels. Beginners in deep learning can generally start by training a supervised deep neural network on a modestly sized dataset, typically containing hundreds to a few thousand well-labelled examples.A clean labelled dataset refers to data in which the labels are accurate and consistent, without mis labelled or ambiguous examples.
深度学习算法可分为监督学习、无监督学习和强化学习。监督学习是最常见的深度学习类型。在监督学习中,模型通过带标签的数据集进行训练,这意味着每个训练样本(例如一张照片)都配有输出标签或真实标签(例如"猫"或"狗")。模型根据输入数据做出预测或决策,并通过真实标签学习正确预测的模式。深度学习初学者通常可以从在中等规模数据集上训练监督式深度神经网络开始,这类数据集通常包含数百至数千个标注良好的样本。干净的标注数据集是指标签准确一致、没有错误标注或模糊样本的数据集。
By contrast, unsupervised learning is a method in which the model is trained toidentify patterns ina dataset without any explicit labels or annotations. It is commonly used in scenarios such as clustering, anomaly detection and association tasks,inwhich the model determines the inherent structure ofthe data to draw inferences°.Self-supervised learning (SSL) is a specific approach within unsupervised learning that involves training a deep neural network ona dataset in which labels are not provided in a traditional sense.Instead,inSSL,the model generates its own labels through a pseudo-task'. For example, the model might be tasked with reconstructing an image from a distorted version or predicting a missing word in a sentence. Nowadays, SSL is commonly used as thefirst stepto'pre-train'deep neural networks for subsequent 'fine-tuning' with supervised learning&,9.
相比之下,无监督学习 (Unsupervised Learning) 是一种模型在没有明确标签或标注的情况下识别数据集中模式的方法。它通常用于聚类 (clustering) 、异常检测 (anomaly detection) 和关联任务 (association tasks) 等场景,模型通过确定数据的内在结构来得出推断 [20]。自监督学习 (Self-supervised Learning, SSL) 是无监督学习中的一种特定方法,涉及在传统意义上未提供标签的数据集上训练深度神经网络。在 SSL 中,模型通过伪任务 (pseudo-task) 生成自己的标签。例如,模型可能被要求从失真版本重建图像,或预测句子中缺失的单词。如今,SSL 通常作为预训练 (pre-train) 深度神经网络的第一步,以便后续通过监督学习进行微调 (fine-tuning) [8,9]。
The third class of deep learning approaches is reinforcement learning. A reinforcement learning system involves agents, entities or systems that make decisions by interacting with their surroundings to achieve a goal. Over time, the agent learns the optimal behaviour based on feedback from the environment. This feedback is in the form of numerical rewards for actions that lead to successful outcomes and penalties for those that do not. The goal is for the agent to learn a policy that maximizes the cumulative reward overtime.Reinforcement learning can automate rule-based procedures such as computer games10 or drone navigation. Reinforcement learning is applied in cancer research in some specific areas, such as for finding optimal policies for personalized cancer screening? or designing clinical trials.
深度学习的第三类是强化学习 (Reinforcement Learning)。强化学习系统包含智能体 (Agent),即通过与周围环境交互来制定决策以实现目标的实体或系统。随着时间的推移,智能体会根据环境反馈学习最优行为模式。这种反馈以数值形式呈现:导致成功结果的动作获得奖励,而未达标的动作则受到惩罚。其目标是让智能体学会能随时间累积最大奖励的策略。强化学习可自动化基于规则的程序,例如电脑游戏[10]或无人机导航。在癌症研究领域,强化学习已应用于某些特定场景,例如寻找个性化癌症筛查的最优策略,或设计临床试验方案。
Neural network architectures
神经网络架构
Deep learning can effectively handle unstructured data such as images and text. Unstructured data refer to information that lacks a specific schema and does not follow conventional data models1². The discipline of automatically analysing images with computers is called computer vision, whereas the discipline dedicated to analysing and interpreting human language intext form is known as NLP. Throughout most ofthe 2010s,convolutional neural networks(CNNs)have been at the core of advancements in computer vision'3, whereas in the field of NLP, long short-term memory (LSTM) networks and related architectures have been widely usedl. LSTM networks, known for their ability to handle sequential data, were a substantial improvement over earlier models due to their capacity to remember information over long sequences. Despite their effectiveness, LSTM networks sometimes struggled with highly complex or extended sequences 15. This limitation has led to the exploration and adoption of alternative architectures in the 2020s, such as the transformer neural networks, also called transformers, which addresses some of these challenges more effectively in certain NLP tasks16. Interestingly, transformers can also be applied to images and even outperform CNNs at many medical image analysis tasks17,18. Transformers can capture long-range dependencies and global context in the images, whereas CNNs are by design constrained to more
深度学习能有效处理图像和文本等非结构化数据。非结构化数据指缺乏特定模式且不遵循传统数据模型的信息[1][2]。利用计算机自动分析图像的学科称为计算机视觉,而专门分析和解释文本形式人类语言的学科则被称为自然语言处理 (NLP)。在整个2010年代的大部分时间里,卷积神经网络 (CNN) 一直是计算机视觉领域进步的核心[3],而在自然语言处理领域,长短期记忆 (LSTM) 网络及相关架构被广泛使用。LSTM 网络以其处理序列数据的能力而闻名,由于能够记忆长序列信息,相较早期模型有显著改进。尽管效果显著,但 LSTM 网络有时难以处理高度复杂或超长序列[5]。这一局限性促使人们在2020年代开始探索采用替代架构,例如 Transformer 神经网络(也称 Transformer),该架构在某些自然语言处理任务中能更有效地应对这些挑战[16]。值得注意的是,Transformer 也可应用于图像处理,并在许多医学图像分析任务中表现优于卷积神经网络[17][18]。Transformer 能捕捉图像中的长程依赖关系和全局上下文信息,而卷积神经网络在设计上受限于局部感受野。
Review article
综述文章
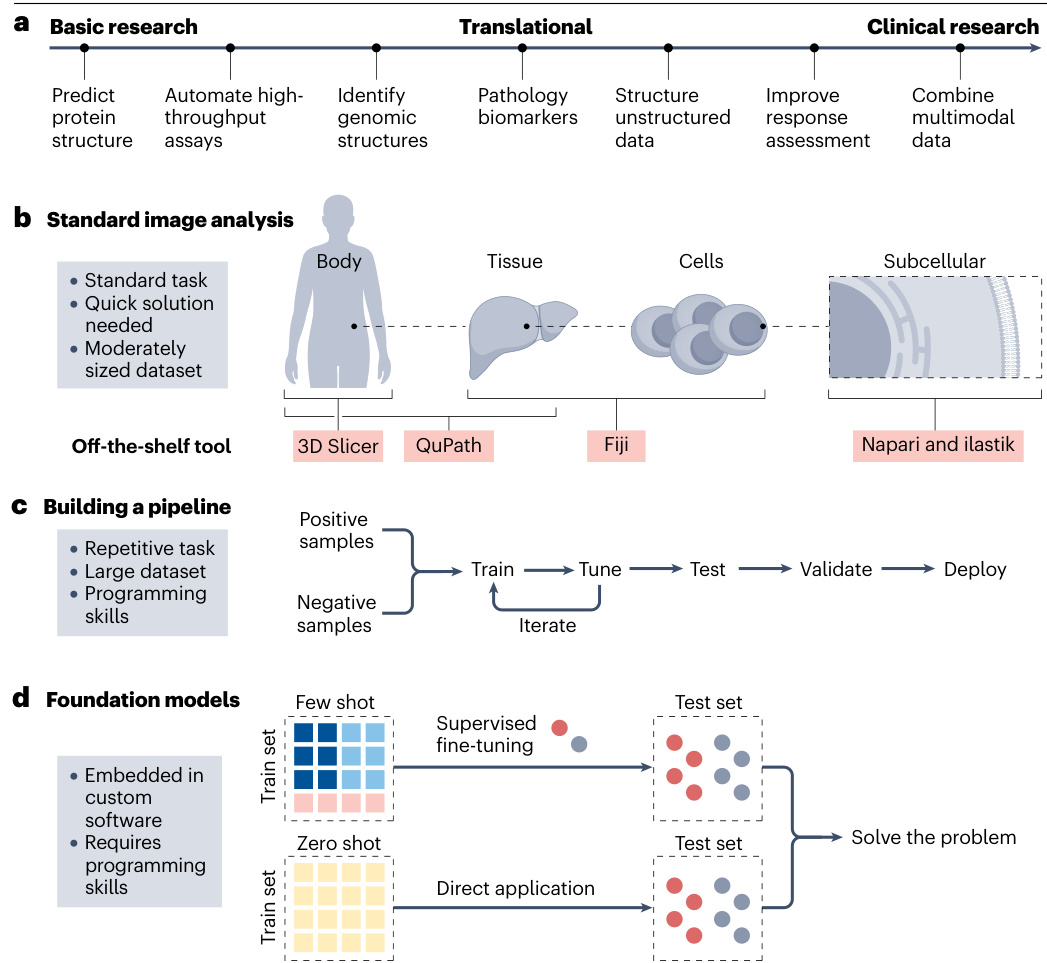
Fig.1|Al workflows in cancer research.
图 1|癌症研究中的AI工作流程。
a,Applications of deep learning connect basic, translational and clinical research.In artificial intelligence (Al)-based image processing,three approaches prevail(asshowninb-d).b,Standard image analysis tasks such as cell counting or segmenting regions ofinterest(ROls)often require swift solutions with off-the-shelf tools.Numerous open-source tools are available for accomplishing these tasks.For example,ilastik is well-suited for subcellular analysis,whereas QuPath and 3D Slicer are commonly used for tissue-leveland whole-body imaging,respectively.Fiji,ImageJ and Cell Profiler are versatile tools that can handle a wide range of image analysis tasks across different scales.A selection of these tools is presented in Supplementary Table1.c,Another categoryof image analysis tasks involves those that require an iterative process of training and fine-tuning on extensive datasets.These tools are typically very task-specific and are designed to address one particular question.Constructing,training and validating these pipelines requires programming skills. d, The future of image analysis is trending towards foundation models,which are trained on extensive, heterogeneous datasets and can be finetuned on numerous tasks.Inzero-shot learning,the model is applied directly to the test set without any task-specific training,whereas in few-shot learning, the model is provided with a small number of labelled examples to adapt to the specific task.
a, 深度学习应用连接基础研究、转化研究和临床研究。在基于人工智能 (AI) 的图像处理领域,主要存在三种方法 (如 b-d 所示)。b, 标准图像分析任务 (如细胞计数或感兴趣区域 (ROI) 分割) 通常需要利用现成工具快速解决。现有众多开源工具可完成这些任务,例如 ilastik 适用于亚细胞分析,而 QuPath 和 3D Slicer 分别常用于组织水平和全身成像。Fiji、ImageJ 和 Cell Profiler 是多功能工具,可处理不同尺度的各种图像分析任务。部分工具选列见补充表 1。c, 另一类图像分析任务需要对大规模数据集进行迭代式训练和微调。这类工具通常具有高度任务特异性,旨在解决特定问题。构建、训练和验证这些流程需要编程技能。d, 图像分析的未来趋势是基础模型,这些模型在广泛异构数据集上训练而成,可针对众多任务进行微调。在零样本学习中,模型直接应用于测试集而无需任何任务特定训练;而在少样本学习中,模型通过少量标注示例来适应特定任务。
localized patterns19. As of 2024, transformers represent the state of the art in both image and language processing tasks, showcasing their versatility and capability in handling diverse and complex data types. In particular, transformers coupled with SSL have catalysed a movement towards so-called foundation models,which is discussed further below (Figs.1b-d and 2a,b).
localized patterns19. 截至2024年,Transformer架构已成为图像与语言处理任务的最先进技术,展现了其处理多样复杂数据类型的通用性与强大能力。特别是结合自监督学习 (SSL) 的Transformer架构催生了所谓基础模型 (foundation models) 的发展浪潮,下文将对此展开进一步探讨 (图1b-d 和图2a,b)。
Uses of deep learning in cancer research
深度学习在癌症研究中的应用
Deep learning has a wide array of applications in cancer research and could improve the productivity of researchers (Box 2). Its capability to analyse unstructured data is applicable for the analysis of results from many experimental assays, such as microscopic images, as well as text from various sources in research and clinical routines. Deep learning also effectively handles other complex data types, including genomic information. In these settings, deep learning is utilized in twomain ways.Thefirstis through user-friendly tools that require no programming skills, offering graphical interfaces for ease of use. For example,software such as Qu Path ori last ik provide intuitive inter- faces for analysing microscopy images and can be extended with deep learning models for tasks such as cell segmentation or classification. The second, more advanced method involves using programming languages such as Python to interact with deep learning architectures, such as CNNs or transformers, through scripts. This approach provides greater flexibility but necessitates a basic understanding of programming(Fig.1b,c).
深度学习在癌症研究中具有广泛的应用,并能提升研究人员的工作效率 (Box 2) 。其分析非结构化数据的能力适用于处理多种实验检测结果,例如显微图像,以及研究和临床流程中来自不同来源的文本。深度学习还能有效处理其他复杂数据类型,包括基因组信息。在这些场景中,深度学习主要通过两种方式应用:第一种是通过无需编程技能的友好型工具,提供图形界面以方便使用。例如,Qu Path 或 i last ik 等软件为分析显微镜图像提供了直观界面,并可通过集成深度学习模型来执行细胞分割或分类等任务。第二种更进阶的方法涉及使用 Python语言 等编程语言,通过脚本与卷积神经网络 (CNN) 或 Transformer 等深度学习架构交互。这种方法灵活性更高,但需要掌握基础的编程知识 (图 1b,c) 。
Deep learning algorithms are also used to analyse medical imaging data, such as detecting tumours in MRIor CT scans with precision comparable to that of experienced radiologists 20.21 This technology also assists in identifying subtle patterns in genetic data??, resulting in the understanding of the genetic origins of a cancer. In drug discovery, deep learning can help in screening potential compounds more efficiently by analysing both in silico (computational) and in vitro (experimental) data, speeding up the process of finding new cancer treatments 23. Furthermore, deep learning is used in his to pathology to analyse tissue samples, distinguishing between benign and malignant cells with remarkable accuracy24, or extracting clinically usable biomarkers directly from image data2.
深度学习算法也被用于分析医学影像数据,例如通过核磁共振成像或计算机断层扫描检测肿瘤,其精准度可与经验丰富的放射科医师相媲美 [20][21]。这项技术还有助于识别基因数据中的细微模式,从而理解癌症的遗传起源。在药物发现领域,深度学习通过分析计算机模拟数据和体外实验数据,能更高效地筛选潜在化合物,加速新型癌症疗法的研发进程 [23]。此外,深度学习应用于数字病理学领域时,可分析组织样本以极高准确度区分良恶性细胞 [24],或直接从影像数据中提取具有临床价值的生物标志物 [2]。
Al for biomedical image analysis
生物医学图像分析人工智能
The breakthroughs in deep learning in the 2010s,such asthe development of CNNs and the availability of large-scale annotated datasets, broadened the applicability of computer-based image analysis to more complex tasks(Fig.1c).Forexample,classical machine learning methods before the advent of deep learning could detect cells in microscopy images. These approaches were further applied for many downstream applications,such as prognostication in cancer based on enumeration of lymphocytes in histology slides25. These early computer vision tools inthe1990s andearly 2000s reliedon handcrafted features, such as edge detection, texture analysis and colour-based segmentation, which required domain expertise and were often limited in their genera liz ability across different datasets.Although simple cell detection has been improved and made more general iz able with deep learning methods,as compared with pre-deep learning tools, such as the commonly used tool StarDist26, deep learning is not strictly necessary to
20世纪10年代深度学习的突破,例如卷积神经网络 (CNN) 的发展和大规模标注数据集的可用性,将基于计算机的图像分析的适用性扩展到更复杂的任务 (图 1c) 。例如,在深度学习出现之前,经典机器学习方法可以检测显微镜图像中的细胞。这些方法被进一步应用于许多下游应用,例如基于组织学切片中淋巴细胞计数的癌症预后预测25。20世纪90年代和21世纪初的这些早期计算机视觉工具依赖于手工制作的特征,例如边缘检测、纹理分析和基于颜色的分割,这需要领域专业知识,并且在不同数据集间的泛化能力通常有限。虽然与深度学习前的工具 (例如常用工具 StarDist26) 相比,简单细胞检测已通过深度学习方法得到改进并变得更可泛化,但严格来说深度学习并非必需。
Review article
综述文章
Box1
Evolution of Al from theory to practical application
人工智能从理论到实际应用的演进
The field of artificial intelligence (Al) is largely regarded as emerging from a conference at Dartmouth College in 1956. Early work was focused on so-called symbolic Al such as rule-based systems for tasks such as playing chess. By the 1960s and 1970s, it became more common to incorporate machine learning algorithms and simplistic artificial neural networks, although these were computationally limited. Machine learning has replaced symbolic Al in virtually all advanced Al applications today, including in cancer research. In the 1980s, the development of the back propagation algorithm enhanced the training efficiency of artificial neural networks. Despite these advancements, Al research faced a winter period in the late 1980s and 1990s due to unfulfilled expectations and subsequent funding cuts. The focus then shifted towards machine learning techniques, such as support vector machines or decision trees, which demonstrated statistically significant performance in various applications. Today, we callthese machine learning methods'classical' machine learning. Nevertheless, artificial neural networks developed further and, finally, in the early 2010s, deep neural networks demonstrated remarkable performance in areas such as image and speech recognition. One of the first real-world applications of neural networks was in postal services. In the 1980s, artificial neural networks were already used to automate the sorting of postal mail by recognizing handwritten letters on envelopes 168. Today, deep learning is widespread, aiding in cancer diagnosis through image analysis, fraud detection in finance and navigation in autonomous vehicles. Al has transitioned from a theoretical discipline to a practical tool with a transformative effect on various industries. Its capabilities now range from basic pattern recognition to advanced neural networks that can learn unsupervised and process natural language in real-time. Whether in smart homes that adapt to us or recommendation algorithms that anticipate our preferences, the value of Al in tackling complex problems has proven to be enormous, and its influence continues to expand.
人工智能 (AI) 领域普遍被认为起源于1956年达特茅斯学院的一次会议。早期研究主要集中于所谓的符号人工智能,例如用于国际象棋等任务的基于规则的系统。到1960年代和1970年代,尽管存在计算能力限制,整合机器学习算法和简化人工神经网络的做法已更为普遍。如今在几乎所有先进AI应用领域(包括癌症研究),机器学习已完全取代符号人工智能。1980年代,反向传播算法的发展提升了人工神经网络的训练效率。尽管取得这些进展,由于预期未能实现及随之而来的资金削减,AI研究在1980年代末至1990年代遭遇寒冬期。研究重心随后转向支持向量机或决策树等机器学习技术,这些技术在各类应用中展现出统计学意义的显著性能。如今我们称这些机器学习方法为"经典"机器学习。然而人工神经网络持续发展,最终在2010年代初,深度神经网络在图像和语音识别等领域展现出卓越性能。神经网络的首个实际应用案例出现在邮政服务领域。1980年代,人工神经网络已通过识别信封上的手写字母来实现邮政自动分拣[168]。当前深度学习技术已广泛应用,通过图像分析辅助癌症诊断、金融欺诈检测以及自动驾驶车辆导航。AI已从理论学科转型为具有变革意义的实用工具,其能力范围涵盖从基础模式识别到能进行无监督学习并实时处理自然语言的先进神经网络。无论是在适配用户需求的智能家居,还是预测用户偏好的推荐算法中,AI在解决复杂问题方面的价值已被证明巨大无比,其影响力仍在持续扩展。
obtain a reasonable detection of cells in microscopy images. However, the situation is different in more complex tasks. In the case of more subtle properties such as biomarker analysis from patient samples, almost all clinically approved systems are based on deep learning22. Many of these biomarkers are based on the simple principle of image classification, which will be discussed below.
在显微镜图像中获得合理的细胞检测。然而,在更复杂的任务中情况有所不同。对于从患者样本中分析生物标志物这类更精细的特性时,几乎所有临床批准的系统都基于深度学习22。这些生物标志物大多基于简单的图像分类原理,下文将对此进行讨论。
Cellular and molecular imaging analysis
细胞与分子成像分析
Cancer researchers often deal with digital images.Inthe case of basic research, these experiments can involve visual assessments, such as visually examining cell cultures for confluency, morphology or growth as a preliminary step or assessing tumour growth in vivo through microscopic methods such as bright field or fluorescence microscopy. Many image analysis tasks in biological research are traditionally performed manually, however this is not only inefficient and error-prone but can also make experiments infeasible if thousands of output images have to be analysed. In general, by using deep learning to quantify experimental readouts, the analysis can be made more objective, reliable and quicker. For instance, in the context of cell detection in phase-contrast microscopy, deep learning can quickly and reliably detect individual cells and classify them as live or dead27. Such analyses are being widely used,for example,through commercial platforms such as the Incucyte Al Cell Health Analysis Software Module (Sartori usA G).
癌症研究人员经常处理数字图像。在基础研究领域,这些实验可能涉及视觉评估,例如通过目测检查细胞培养物的汇合度、形态或生长情况作为初步步骤,或通过明场或荧光显微镜等显微方法评估体内肿瘤生长。生物学研究中的许多图像分析任务传统上依赖人工完成,但这种方式不仅效率低下且容易出错,当需要分析数千张输出图像时甚至可能导致实验无法实施。总体而言,通过使用深度学习技术量化实验读数,可以使分析过程更具客观性、可靠性并提升效率。例如在相差显微镜下的细胞检测场景中,深度学习能够快速可靠地识别单个细胞并将其分类为存活或死亡细胞[27]。此类分析正得到广泛应用,例如通过商业化平台如Incucyte AI细胞健康分析软件模块(Sartorius AG)实现。
Open-source Al solutions for microscopy image analysis. In general,commercially available software utilizing deep learning forbasic science research is only available for common and standardized tasks, such as cell counting in phase-contrast images. The open-source community, however, has made dozens of deep learning methods available for the analysis of microscopy images. For example, QuPath28 is a common software for viewing gigapixel microscopy images, in which a single image file contains multiple gigabytes of compressed data, and it enables access to popular deep learning models for cell detection such as 'stardist'26 without requiring any programming skills. Similarly, ImageJ",ImageJ2 (ref. 30) and Fiji²1 are the standard tools for many image viewing and analysis tasks in biology, including viewing multichannel, multidimensional images, even in more uncommon file formats such as CZI or MRxS, via the integration with Bio-Formats32 (Supplementary Table 2).Onthese platforms,pre-trained deeplearning models can be run through various open-source platforms and plugins, including deep lm age J 33. Some pre-trained models for cell segmentation, nucleus segmentation or more specialized tasks such as segmentation of mitochondria in electron microscopy images are available through a repository or collection of pre-trained models, termed model'zoos' (for example, available at Biolmage.io).
用于显微镜图像分析的开源AI解决方案。总的来说,利用深度学习进行基础科学研究的商业软件仅适用于常见和标准化任务,例如相差图像中的细胞计数。然而,开源社区已经提供了数十种深度学习方法用于显微镜图像分析。例如,QuPath28是查看千兆像素显微镜图像的常用软件,其中单个图像文件包含多个千兆字节的压缩数据,并且无需任何编程技能即可访问流行的深度学习模型进行细胞检测,例如'stardist'26。类似地,ImageJ"、ImageJ2(参考文献30)和Fiji²1是生物学中许多图像查看和分析任务的标准工具,包括查看多通道、多维图像,甚至通过Bio-Formats32的集成(补充表2)查看更不常见的文件格式,如CZI或MRxS。在这些平台上,预训练的深度学习模型可以通过各种开源平台和插件运行,包括deep lm age J 33。一些用于细胞分割、细胞核分割或更专业任务(如电子显微镜图像中线粒体分割)的预训练模型可通过存储库或预训练模型集合(称为模型"动物园")获得(例如,可在Biolmage.io上获得)。
For many specialized niche applications, no off-the-shelf model or platform is available.In these cases,researchers are best advised to build their own software based on deep learning and redistribute it to other researchers under an open-source license. For example, deep learning has been successfully used in research software pipelines by several academic research groups for the purpose of evaluating tumour organoids in bright field microscopy 34,35. These research groups have made their software available to other researchers under open-source licenses, enabling wider adoption and collaboration. Furthermore, some advanced biological imaging techniques, such as reconstructing high-resolution fluorescence images from low-resolution or noisy data, cannot be performed efficiently without deep learning3. Although ImageJ can perform basic image reconstruction tasks such as tiling, deep learning-based methods can substantially improve the quality of the reconstructed images by learning to remove noise, enhance resolution and infer missing information from the available data.Often, research groups must develop advanced computational methods,such as custom deep learning architectures or novel training strategies,to enable them to solve specific image analysis needs for which nogeneral solution is available. These custom models can circumvent the limitations of traditional image reconstruction techniques by learning to exploit patterns and relationships within the data that are not easily captured by handcrafted algorithms.
对于许多专业细分领域的应用场景,市面上并没有现成的模型或平台可供使用。在这种情况下,研究人员的最佳选择是基于深度学习技术自主开发软件,并通过开源许可协议分享给其他研究者。例如,多个学术研究团队已成功将深度学习应用于研究软件流程中,用于明场显微镜下的肿瘤类器官评估 [34,35] 。这些研究团队通过开源许可将软件开放给其他研究者,促进了更广泛的采用与合作。此外,某些先进的生物成像技术(如从低分辨率或含噪数据重建高分辨率荧光图像)若没有深度学习将无法有效实现 [3] 。尽管 ImageJ 能够执行基本的图像重建任务(如图像拼接),但基于深度学习的方法通过学习去除噪声、增强分辨率以及从现有数据推断缺失信息,能显著提升重建图像的质量。研究团队通常需要开发先进的计算方法(例如定制化的深度学习架构或创新的训练策略),以满足特定图像分析需求,因为这类需求往往缺乏通用解决方案。这些定制模型通过学习数据中难以通过手工算法捕捉的模式与关联,能够规避传统图像重建技术的局限性。
His to pathology
组织病理学
Capabilities of deep learning in pathology image analysis. In basic and translational cancer research, tumour tissue from patients or animal models is analysed by histology 17 to diagnose and evaluate tumours.The his to logical morphology of a tumour represents the
深度学习在病理图像分析中的能力。在基础与转化癌症研究中,来自患者或动物模型的肿瘤组织通过组织学17进行分析以诊断和评估肿瘤。肿瘤的组织学形态代表了
Review article
综述文章
result of a plethora of molecular processes in the genome ofatumour, epigenome and environment. Because ofthis,it is essential to digitize these slides into high-resolution, gigapixel images. The challenge in analysing these images arises from their immense size combined with their detailed content. This complexity often exceedsthe capabilities of standard microscopy image analysis tools, such as ImageJ or Fiji29.31 particularly when dealing with a large volume of data. Although ImageJ and Fiji provide powerful tools for image processing and analysis, including the ability to create macros and scripts for automated workflows, they are not optimized for handling gigapixel images or large datasets. Other software, such as QuPath, is optimized for handling such gigapixel images, but there are many other ways to use deep learning in pathology image analysis, which we describe below.
肿瘤基因组、表观基因组和环境中大量分子过程作用的结果。正因如此,必须将这些玻片数字化为高分辨率的十亿像素级图像。分析这些图像的挑战在于其巨大的尺寸与精细内容相结合。这种复杂性往往超出了标准显微镜图像分析工具(如ImageJ或Fiji [29.31])的处理能力,特别是在处理海量数据时。尽管ImageJ和Fiji提供了强大的图像处理与分析工具(包括创建宏和脚本实现自动化工作流程),但它们并未针对十亿像素级图像或大型数据集进行优化。其他软件(如QuPath)虽针对此类十亿像素级图像进行了优化,但在病理图像分析中应用深度学习还有许多其他方法,我们将在下文详述。
Computational pathology. Although digital pathology involves obtaining, storing and viewing these images, the analysis is referred to as computational pathology. From atechnical point of view, before the broader application of deep learning models, image analysis pipelines were composed of successive, highly engineered steps,tailored to specific types of analysis. For example, a machine learning model would detect cells, after which a subsequent model would classify cells into distinct cell types, and a final model would predict patient prognosis from these measurements?5. However, it is difficult to build such multistep pipelines with classical machine learning methods because of the complexity of integrating the multiple processing stages. The complexities of these tasks have resulted in the growing adoption of deep learning-based end-to-end models as an alternative approach17, as they consist of fewer steps and rely on less explicitly defined expert knowledge. They use deep neural networks and can learn directly from raw data, to generate the output without the need for different intermediate steps.
计算病理学。虽然数字病理学涉及获取、存储和查看这些图像,但其分析过程被称为计算病理学。从技术角度来看,在深度学习模型广泛应用之前,图像分析流程由一系列连续的、高度工程化的步骤组成,这些步骤针对特定类型的分析进行了定制。例如,机器学习模型会先检测细胞,随后另一个模型将细胞分类为不同细胞类型,最终模型根据这些测量结果预测患者预后[5]。然而,由于需要整合多个处理阶段,使用经典机器学习方法构建这种多步骤流程十分困难。这些任务的复杂性促使人们越来越多地采用基于深度学习的端到端模型作为替代方案[17],因为这类模型步骤更少,且依赖较少明确定义的专家知识。它们使用深度神经网络,能够直接从原始数据中学习并生成输出,无需不同的中间步骤。
Tasks in computational pathology. Computational pathology can be applied to solve two types oftasks. One type oftask includes recapitulating human evaluation of images, such as counting cells or measuring tissue size. According to Echle37, basic'tasks are intended to simplify routine workflows that are currently performed solely by pathologists. These basic tasks can, in principle, be performed by humans, but are time-consuming and not scalable if done by hand38. By contrast, advanced tasks, as per the previously established definition 22.37, comprise the prediction of more high-level properties directly from image data, such as predicting genetic alterations directly from haematoxylin and eosin (H&E)-stained tissue37. Some genetic alterations, such as micro satellite instability (Msl) in colorectal cancer, are linked
计算病理学的任务。计算病理学可用于解决两类任务。一类任务包括重现人类对图像的评估,例如细胞计数或组织尺寸测量。根据Echle[37]的观点,基础任务旨在简化目前仅由病理学家执行的常规工作流程。这些基础任务原则上可由人工完成,但耗时且难以规模化操作[38]。相比之下,按照先前确立的定义[22,37],高级任务包含直接从图像数据预测更高层次的特性,例如直接从苏木精-伊红(H&E)染色组织预测基因突变[37]。某些基因突变(如结直肠癌中的微卫星不稳定性(MSI))与
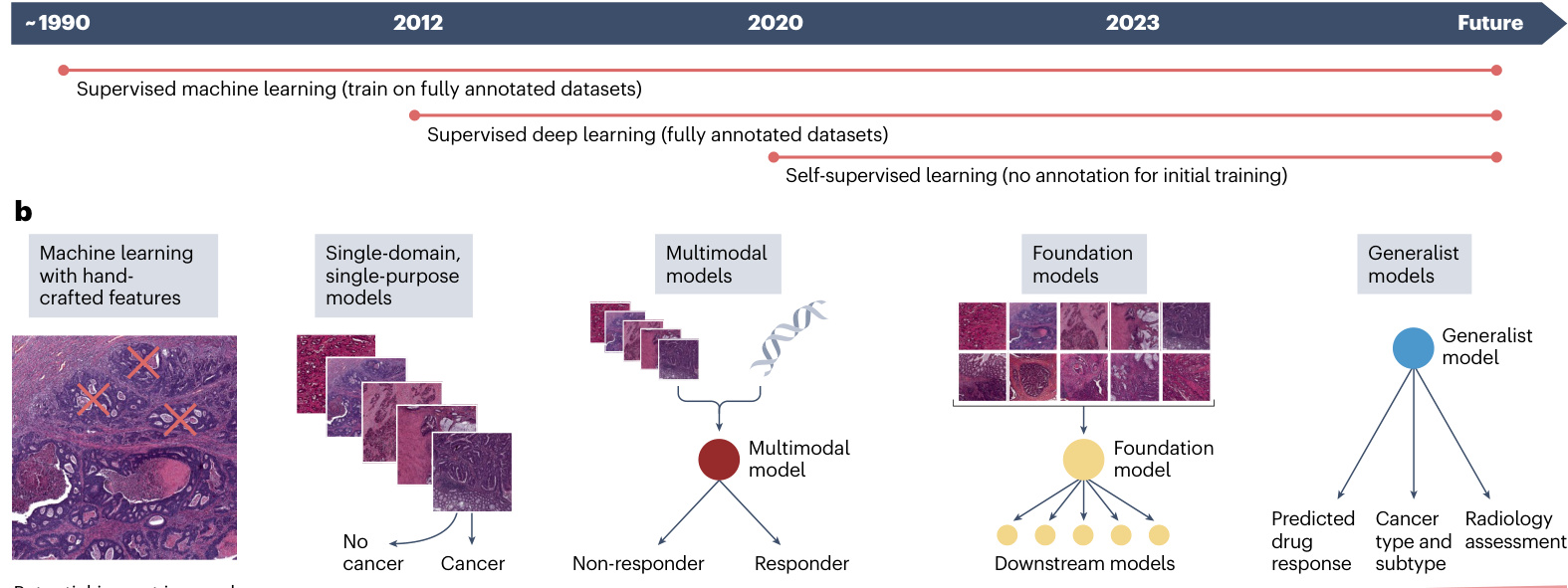
a Fig.2|Development from simple,specialized,shallow models to deep, multimodal,generalist models for computer vision.a, The progress of artificial intelligence (Al) techniques in medical imaging started to be widely used with supervised learning using machine learning models with handcrafted features in the early 2o00s. These models relied on domain expertise to manually extract relevant features from images,which were then used totrainthe models on labelled datasets.Around 2012, supervised deep learning emerged, in which Al models,particularly convolutional neural networks (CNNs),were trained on large labelled datasets to automatically learn hierarchical features directly from raw image data.This approach markedly improved the performance and general iz ability of Al models in medical imaging tasks.Inthe early2020s, many research groups started to use the emerging self-supervised learning methods,which enable models to learn meaningful patterns from unlabelled data by predicting properties of the data itself, without relying on external labels.b, In the initial stages of Al application to cancer research,the approach
图 2 | 从简单、专用、浅层模型到深度多模态通用模型的计算机视觉发展历程。a, 人工智能 (AI) 技术在医学影像中的进展始于 2000 年代初采用具有手工特征特征的机器学习模型进行监督学习。这些模型依赖领域专业知识从图像中手动提取相关特征,然后利用标记数据集训练模型。约 2012 年,监督深度学习兴起,AI 模型(特别是卷积神经网络 (CNN))通过大型标记数据集训练,直接从原始图像数据中自动学习层次化特征。该方法显著提升了 AI 模型在医学影像任务中的性能与泛化能力。2020 年代初,许多研究团队开始采用新兴的自监督学习方法,使模型能够通过预测数据自身属性(无需依赖外部标签)从未标记数据中学习有意义模式。b, 在 AI 应用于癌症研究的初始阶段,该方法
Potential impact in oncology
肿瘤学领域的潜在影响
was to rely on carefully selected patterns from medical images as a basis for analysis.Overtime,the need for manual features election was eliminated,and models were able to learn essential features directly from the data.Presently, the emphasis is on integrating various data sources through multimodal models, which combine information from different data modalities,such as radiology images,pathology slides,genomic data and clinical records.The envisioned future involves foundation models,which are large-scale,self-supervised models pre-trained on diverse, unlabelled datasets across multiple modalities. These models can be fine-tuned for various downstream tasks with minimal task-specific training data.The ultimate aim of Al application in oncology could be a universal generalist model, a multi-purpose tool with the ability to analyse, interpret and interact with both patients and medical professionals. This generalist model would integrate data from multiple sources, support diagnosis and treatment recommendations, and explain its decisions in a hum z er pre table manner.
过去的方法依赖于从医学图像中精心挑选模式作为分析基础。随着时间推移,不再需要手动特征选择,模型能够直接从数据中学习关键特征。当前的重点是通过多模态模型整合各种数据源,这些模型融合了来自不同数据模态的信息,例如放射影像、病理切片、基因组数据和临床记录。未来的愿景是建立基础模型 (foundation models),这些大规模自监督模型通过跨多模态的多样化未标注数据集进行预训练,只需极少量任务特定训练数据即可微调适应各种下游任务。肿瘤学人工智能应用的终极目标可能是开发通用全能模型,这种多用途工具能够分析、解读数据并与患者及医疗专业人员互动。该全能模型将整合多源数据,支持诊断治疗建议,并以人类可理解的方式解释其决策依据。
Review article
综述文章
Box2
Productivity gains through deep learning
通过深度学习提升生产效率
Beyond direct research applications, deep learning is transforming research workflows by optimizing everyday tasks. Researchers can use artificial intelligence (Al) for administrative tasks, such as drafting standard e-mail responses or creating visual sketches. According to a 2023 report by McKinsey titled The Economic Potential of Generative Al: The Next Productivity Frontier 169, generative Al could affect between $2.6%$ and $4.5%$ of annual revenues in the health-care and pharma sector, translating to an annual additional value between $\cup\mathbb{S}\$60$ billion and $0\mathsf{S}\$110$ billion globally. Similarly, a recent study from the Boston Consulting Group concluded that Al made management consultants significantly more productive by using large language models in routine tasks that involve the generation of text170. Although these are projections not certainties, it could be extrapolated to individual researchers who can become more productive by incorporating Al methods into their daily workflows.
除了直接的研究应用,深度学习正在通过优化日常任务来改变研究工作流程。研究人员可以利用人工智能 (AI) 处理行政事务,例如起草标准电子邮件回复或创建视觉草图。根据麦肯锡2023年发布的报告《生成式AI的经济潜力:下一轮生产力前沿》[169],生成式AI可能影响医疗健康和制药行业年收入的2.6%至4.5%,相当于全球每年新增600亿至1100亿美元价值。同样,波士顿咨询集团近期研究表明,通过在涉及文本生成的常规任务中使用大语言模型,AI使管理咨询师的工作效率显著提升[170]。尽管这些是预测而非确定结论,但可以推断个体研究人员通过将AI方法融入日常工作流程也能获得效率提升。
to known morphological features, which was known even before deep learning39. However, deep learning methods can automatically analyse these morphological features and make quantitative predictions about underlying genetic alterations just by observing routine pathology slides. Deep learning methods have rediscovered known links between the genotype and the phenotype 18,40.41 and could therefore potentially be extended to detect new genotype-phenotype links for many other genetic alterations that have unknown morphological manifestations. Various studies have shown that the mutation status of individual genes42-48, defective DNA repair mechanisms such as $\mathbf{M}\mathbf{S}\mathbf{I}^{18,40,49-51}$ and homologous recombination deficiency*, or tumour mutation burden53-5 are predictable from H&E slides by deep learning. Furthermore, these models have the capability to discern morphological patterns within both epithelial and stromal tumour regions, or to predict the hormone receptor expression status in breast cancer samples without the need for the immuno his to chemistry or immuno fluorescence against the receptors 4256-58. In summary,the application of deeplearning to pathology has demonstrated that the molecular properties of tumours can be identified from basic H&E images.
已知形态学特征与基因型之间的关联在深度学习出现之前就已为人所知39。然而深度学习方法能够自动分析这些形态学特征,仅通过观察常规病理切片即可对潜在基因改变进行定量预测。深度学习方法重新发现了基因型与表型之间的已知联系18,40,41,因此有望扩展到检测许多其他具有未知形态学表现的基因改变的新基因型-表型关联。多项研究表明,单个基因的突变状态42-48、缺陷DNA修复机制如$\mathbf{MSI}^{18,40,49-51}$和同源重组缺陷52,或肿瘤突变负荷53-55,都可以通过深度学习从H&E切片中预测。此外,这些模型能够辨别上皮和间质肿瘤区域内的形态学模式,或在不需针对受体的免疫组织化学或免疫荧光的情况下预测乳腺癌样本中的激素受体表达状态42,56-58。总之,深度学习在病理学中的应用证明,肿瘤的分子特性可以从基础的H&E图像中识别。
Identifying and quantifying biomarkers from images. Computerbased image classification methods can classify or categorize images into distinct groups based on their visual content. A notable application of this is in the field of medical image analysis, particularly in his to pathology where image classification approaches are used to differentiate his to pathological images based on the presence or absence of cancerous cellss. For example, deep learning can be used to identify tumours within H&E-stained his to logical tissue section slides59. The utility of image classification extends to more complex tasks, which have been referred to as advanced computational pathology tasks 223738. An example of such an application is the categorization of his to pathological images to directly predict high-level properties from images, such as molecular alterations 40.44, tumour of origin for patients with cancer of unknown primary6° survival of patients61.62 or patient responses to immuno therapy'3. These advanced computational pathology tasks demonstrate the potential of deep learning to extract clinically relevant biomarkers directly from his to pathological images. By learning to recognize complex patterns and associations within the image data, deep learning models can identify visual features that correlate with specific molecular alterations, tumour types, patient outcomes or treatment responses.Consequently,image classification methods not only assist in diagnostic processes but also contribute to the development of predictive biomarkers,offering valuable insights for personalized treatment strategies°4. In the next sections, we discuss such his to pathology image analysis tasks in cancer research in moredetail.
识别和量化图像中的生物标志物。基于计算机的图像分类方法可以根据图像的视觉内容将其分类或归入不同的组别。一个显著的应用是在医学图像分析领域, 特别是在组织病理学中, 图像分类方法被用于根据癌细胞的存在与否来区分组织病理学图像。例如, 深度学习可用于识别H&E染色的组织学切片中的肿瘤59。图像分类的效用扩展到更复杂的任务, 这些任务被称为高级计算病理学任务223738。此类应用的一个例子是对组织病理学图像进行分类, 以直接从图像预测高级属性, 例如分子改变40.44, 原发灶不明癌症患者的肿瘤起源6°, 患者生存期61.62或患者对免疫疗法的反应'3。这些高级计算病理学任务展示了深度学习直接从组织病理学图像中提取临床相关生物标志物的潜力。通过学习识别图像数据中的复杂模式和关联, 深度学习模型可以识别与特定分子改变, 肿瘤类型, 患者结局或治疗反应相关的视觉特征。因此, 图像分类方法不仅有助于诊断过程, 还有助于预测性生物标志物的开发, 为个性化治疗策略提供有价值的见解°4。在接下来的章节中, 我们将更详细地讨论癌症研究中的此类组织病理学图像分析任务。
Commercially available and custom-built deep learning tools for computational pathology.Easy-to-use software such as QuPath28 offer some deep learning or machine learning-based capabilities for analysis (Supplementary Table 2) including automated tissue segmentation,tumour detection and classification. Through the use of scripts, QuPath also allows users to train custom models to identify specific cell types or quantify biomarkers. In addition, QuPath supports the integration of external machine learning models, enabling users to apply advanced algorithms for complex analyses without training any new models.Although QuPath provides a user-friendly interface for basic computational pathology tasks,most research groups focused on more advanced applications of Alin his to pathology are typically using self-developed pipelines written in the programming language Python. In addition,researchers often develop their own tools for data annotation or visualization ofthe results,but can also use existing tools; in the past 3 years, several multi-purpose open-source software packages for computational pathology incancer research have emerged, many of which have been re-used widely. These include Fast Pathology 65 s, TIA Toolbox 66, $\mathbf{CLAM}^{67}$ and STAMP6s, which are built on top of generalpurpose data-processing and machine learning frameworks, such as PyTorch°, MONA1°, OpenSlide7 and LibVips"2 (Supplementary Table1).Non-computational cancer researchers will find ituseful to first familiarize themselves with Python programming and then complete basic courses on Python-based image processing (available on many online platforms including commercial platforms such as YouTube,Coursera andedx)before delving into the documentation and application of the computational pathology pipeline repositories.
用于计算病理学的商用和定制深度学习工具。诸如QuPath28等易用软件提供部分基于深度学习或机器学习的功能用于分析(附表2),包括自动化组织分割、肿瘤检测与分类。通过脚本使用,QuPath还允许用户训练定制模型以识别特定细胞类型或量化生物标志物。此外,QuPath支持集成外部机器学习模型,使用户无需训练新模型即可应用先进算法进行复杂分析。虽然QuPath为基础计算病理学任务提供了用户友好界面,但大多数专注于人工智能在病理学中更高级应用的研究团队通常使用基于Python语言自行开发的流程。此外,研究人员常自主开发数据标注或结果可视化的工具,但也可使用现有工具;过去三年间,涌现出若干用于癌症研究中计算病理学的多功能开源软件包,其中许多已被广泛复用。这些包括Fast Pathology65、TIA Toolbox66、$\mathbf{CLAM}^{67}$和STAMP68,它们构建于通用数据处理和机器学习框架之上,例如PyTorch69、MONAI70、OpenSlide71和LibVips72(附表1)。非计算领域的癌症研究者会发现,在深入钻研计算病理学流程库的文档和应用之前,先熟悉Python语言编程并完成基于Python语言的图像处理基础课程(可在许多在线平台获取,包括YouTube、Coursera和edX等商业平台)会很有帮助。
Challenges in computational pathology. Computational pathology not only yields practically useful tools but it can also generate new scientific insight via explain ability methods73.Such methods help us to understand which morphological patterns are related to a prediction by the deep learning system74. However, explain able Al methods are still limited, as retrofitting a deep learning pipeline with explain ability capabilities is not trivial. This is because most deep learning models are inherently complex and opaque, with millions of parameters and multiple nonlinear transformations, making it difficult to trace the decisionmaking process from input to output. Adding explain ability to an existing deep learning pipeline requires carefully designed techniques, such as attention mechanisms, saliency maps or concept activation vectors,which can highlight the relevant regions or features in the input data that contribute to the predictions of the model. Developing and integrating these techniques into a production-ready computational pathology workflow is a non-trivial task that requires expertise in both
计算病理学面临的挑战。计算病理学不仅能产生实用工具,还能通过可解释性方法73生成新的科学见解。这类方法帮助我们理解哪些形态学模式与深度学习系统的预测相关74。然而,可解释人工智能方法仍存在局限,因为为深度学习流程添加可解释性功能并非易事。这主要是因为大多数深度学习模型本质上复杂且不透明,具有数百万个参数和多重非线性变换,使得从输入到输出的决策过程难以追溯。为现有深度学习流程添加可解释性需要精心设计的技术,例如注意力机制、显著图或概念激活向量,这些技术可以突出显示输入数据中与模型预测相关的区域或特征。将这些技术开发并整合到可用于实际生产的计算病理学工作流程中,是一项需要双重专业知识的非平凡任务。
Review article
综述文章
deep learning and interpret able Al. Moreover, the real-world clinical adoption of Al-based work flows in cancer his to pathology is hampered by the lack of digital workflows for pathology in North America and Europe7. The regulatory and ethical considerations, integration challenges within existing workflows, and issues of Al interpret ability and trust among health-care professionals further limits this.
深度学习与可解释人工智能。此外,基于人工智能的癌症组织病理学工作流在北美和欧洲的实际临床应用受到病理学数字化工作流缺失的阻碍[7]。监管与伦理考量、现有工作流程中的整合挑战,以及医疗专业人员对人工智能可解释性与信任度的问题进一步限制了其发展。
Radiology
放射学
Clinical radiology for diagnosing,treating and monitoring therapeutic response can include CT, MRl or positron emission tomography (PET) for cancer detection. Unlike his to pathology, radiology produces inherently digital images and is therefore not limited by the digitization process.
临床放射学用于诊断、治疗和监测疗效反应时,可包括CT、MRI或正电子发射断层扫描 (PET) 用于癌症检测。与组织病理学不同,放射学本身生成数字图像,因此不受数字化过程的限制。
The integration of computer vision models into radiology has markedly enhanced the analytical capabilities ofthe field, revolutionizing the way in which researchers analyse and interpret radiological images. This has paved the way for increased diagnostic accuracy, personalized treatment plans and a deeper understanding of cancer. Traditional machine learning models, which often rely on predefined, handcrafted radiomics features, have shown considerable success in identifying patterns and features in imaging data that are not immediately apparent to the human eye. At the same time, deep learning models, which process raw radiology data, offer a more flexible approach by automatically identifying relevant features without the need for manual intervention. This capability is particularly valuable in handling vast amounts of data, in which deep learning models can uncover complex patterns and relationships within the images. Attempts to inject handcrafted prior knowledge into any machine learning systems are less effective than hypothesis-free or data-driven deep learning approaches, which tend to excel as more data are used, thereby ultimately outperforming handcrafted pipelines. In other words, as the amount of available data increases, deep learning models that automatically learn relevant features from the data tend to surpass the performance of models that rely on manually designed features based on prior knowledge. In radiology, the vast amounts of data lead deep learning to typically outperform classical machine learning methods such as handcrafted radiomics7.7. However, in some scenarios, classical machine learning can offer advantages over deep learning. In situations with limited data(onlyafew dozens to 1 o 0 cases are available),the high capacity of deep learning modelsto learn from data may lead to over fitting, and thus machine learning models using simpler'features'such as tumour sizeand shapecan be more reliable. In general, it is important to choose the appropriate methods for the problem, and only a deep understanding ofthe methodology principles and the data at hand can enable such appropriate decisions.
将计算机视觉模型整合到放射学领域显著增强了该领域的分析能力,彻底改变了研究人员分析和解读放射学图像的方式。这为提高诊断准确性、制定个性化治疗方案以及更深入地了解癌症铺平了道路。传统机器学习模型通常依赖预定义的手工设计放射组学特征,在识别成像数据中肉眼难以直接观察到的模式和特征方面已取得显著成功。与此同时,处理原始放射学数据的深度学习模型通过自动识别相关特征而无需人工干预,提供了更灵活的方法。这种能力在处理海量数据时尤其宝贵,深度学习模型能够从中发掘图像内复杂的模式与关联。相较于依赖假设或无监督的数据驱动深度学习方法,试图将人工先验知识注入任何机器学习系统的效果往往较差——随着数据量增加,深度学习方法往往表现更优,最终超越人工设计的流程。换言之,随着可用数据量的增长,从数据中自动学习相关特征的深度学习模型往往超越依赖基于先验知识手动设计特征的模型性能。在放射学中,海量数据使得深度学习通常优于传统机器学习方法(例如手工放射组学)。但在某些场景下,经典机器学习仍能展现优势。当数据有限(仅几十至百例病例)时,深度学习模型强大的数据学习能力可能导致过拟合,此时使用更简单特征(如肿瘤尺寸和形状)的机器学习模型反而更可靠。总体而言,根据具体问题选择合适方法至关重要,只有深入理解方法论原理和现有数据特性才能做出恰当决策。
Applying Al in radiology: key considerations and approaches. When applying machine learning or deep learning models to medical imaging, a crucial step is preprocessing the data to reduce variability arising from diverse acquisition protocols within the same imaging modality. This involves standardizing image parameters such as slice thickness and voxel size, reducing noise and normalizing signal intensity7. Subsequently, in-depth information from radiological images can be extracted using predefined algorithms that compute standardized radiomics features79. These are handcrafted parameters, also known as engineered radiomics features, extracted from medical images using mathematical algorithms designed to quantify particular aspects of the image, such as signal intensity, texture and shape798o. These provide valuable information about voxel density or signal intensity (depending on the image modality) and also offer insights into the size and shape of the region of interest (ROl). Among the numerous algorithms for extracting handcrafted radiomics from medical images, Py radio mic s is a widely used Python package*°. To use it, a user must first install Py radio mic s and import the necessary libraries into your Python notebook. Next, set up the feature extractor to meet your specific needs,including image preprocessing and feature selection options. After loading the medical image and its associated segmentation mask, which outlines the ROl, the user can proceed to extract radiomics features from the ROl. This customizable process allows for tailoring to your research requirements. Using handcrafted radiomics features, which are usually coupled with classical machine learning models (that is, those utilized before the advent of deep learning architectures, such as logistic regression, support vector machines(SvMs)or random forest)can be trained to test various hypotheses. However, it is important to remember that applying handcrafted radiomics requires meticulous image annotation, such as delineating the R Ol corresponding to tumours in cancer studies. This delineation process should be and is typically performed manually by expert radiologists.Although some software,suchas3D Slicer? or ITK-SNAP?2, provide tools to facilitate this process,it remains a highly time-consuming task.
人工智能在放射学中的应用: 关键考量与方法
将机器学习或深度学习模型应用于医学影像时,关键步骤是通过预处理减少同一成像模态中不同采集协议带来的变异性。这包括标准化图像参数 (如层厚和体素尺寸) 、降噪和信号强度归一化7。随后,可使用预定义算法计算标准化放射组学特征79,从放射影像中提取深度信息。这些通过数学算法从医学影像中提取的手工参数 (亦称人工设计的放射组学特征) ,可量化图像的特定方面,如信号强度、纹理和形状7980。这些特征不仅提供关于体素密度或信号强度 (取决于成像模态) 的宝贵信息,还能揭示感兴趣区域 (ROI) 的大小和形态特征。在众多从医学影像提取手工放射组学的算法中, PyRadiomics 是广泛使用的 Python 语言工具包*°。使用时需先安装 PyRadiomics 并将必要库导入 Python 笔记本,接着根据具体需求配置特征提取器 (包括图像预处理和特征选择选项) 。加载医学影像及其对应的勾画 ROI 的分割掩模后,即可从 ROI 提取放射组学特征。这种可定制流程能有效适配研究需求。
手工放射组学特征通常与经典机器学习模型 (即深度学习架构出现前使用的模型,如逻辑回归、支持向量机或随机森林) 结合,可训练模型验证各种假设。但需注意,应用手工放射组学需要精细的图像标注,例如在癌症研究中勾画对应肿瘤的 ROI。该勾画过程应当且通常由专业放射科医师手动完成。尽管 3D Slicer? 或 ITK-SNAP?2 等软件提供了辅助工具,这仍是极其耗时的任务。
Deep learning applications for radiology. Alternatively, neural networks offer a powerful approach in medical image analysis. Although they can also be trained with data from pre-delineated tumours, they excel at processing whole images. Training neural networks with whole image data can obviatethe labour-intensive process of manual delineation of areas of interest. This also allows the model to learn patterns not only from the tumour but also from the rest of the anatomical structures captured in medical scans. Moreover, unlike traditional methods that rely on predefined handcrafted features, deep learning models can automatically learn relevant features directly from the image data62.83.84. This has a key role in advancing image quality while reducing acquisition timess5.86 Recently, deep learning in radiology has become more robust through large-scale training and is now capable of precisely delineating every organ in the body from whole-body scans, setting a new standard in medical imaging?7. Currently several computer-aided systems are being used to enhance radiology imaging analysis,for instance, as medical devicesfor improving the detection oflung cancer (such as aview LCS, syngo.CT Lung CAD,Samsung Auto Lung Nodule Detection and InferRead CD Lung) and breast cancer (such as breasts cape v1.0 and M ammo Screen in screening programmes and brain tumour diagnosis8. These tumour screening applications of Al-enabled radiology image analysis have also been validated in largescale prospective clinical trials89. Exceeding this, machine and deep learning models have also shown great promise in identifying subtle cancer patterns in radiology imaging, which are associated with specific mutations and the likelihood of responding to different treatments, in both pre clinical and clinical settings90-94. As pre clinical experiments often involve smaller datasets, usually just a few dozen mice compared with hundreds of patients inclinical trials,some applications consider each voxel of the image as input, which helps to mitigate the challenges posed bylimited datafor deep learning models"s. This voxel-wise approach allows for a more detailed analysis by treating each voxel as a separate data point, potentially increasing the sensitivity to subtle features relevant to the study. This method contrasts with using the full image as a source of information. Analysing whole images allows for the integration of contextual and spatial information. Again, having an
深度学习在放射学中的应用。或者,神经网络为医学影像分析提供了强大方法。虽然它们也能通过预勾画肿瘤数据进行训练,但其优势在于处理完整图像。使用完整图像数据训练神经网络可避免耗时费力的人工感兴趣区域勾画过程。这使得模型不仅能够从肿瘤中学习模式,还能从医学扫描捕获的其余解剖结构中学习。此外,与传统依赖预定义手工特征的方法不同,深度学习模型能直接从图像数据中自动学习相关特征[62][83][84]。这在提升图像质量同时缩短采集时间方面具有关键作用[85][86]。近年来,放射学中的深度学习通过大规模训练变得更为稳健,现已能够从全身扫描中精确勾画体内每个器官,为医学影像设立了新标准[87]。目前已有若干计算机辅助系统用于增强放射影像分析,例如作为医疗器械用于提升肺癌检测(如aView LCS、syngo.CT Lung CAD、Samsung Auto Lung Nodule Detection和InferRead CD Lung)和乳腺癌检测(如筛查项目中的breasts cape v1.0和Mammo Screen)以及脑肿瘤诊断[88]。这些基于人工智能的放射影像分析在肿瘤筛查中的应用已通过大规模前瞻性临床试验验证[89]。不仅如此,机器和深度学习模型在识别放射影像中与特定突变及不同治疗响应可能性相关的细微癌症模式方面也展现出巨大潜力,涵盖临床前和临床环境[90-94]。由于临床前实验通常涉及较小数据集(通常仅数十只小鼠,而临床试验有数百名患者),某些应用将图像的每个体素作为输入,这有助于缓解深度学习模型面临的数据有限挑战[95]。这种逐体素方法通过将每个体素视为独立数据点,允许进行更精细分析,可能提高对研究中相关细微特征的敏感性。该方法与使用完整图像作为信息源形成对比。分析完整图像可实现上下文和空间信息的整合。
Review article
综述文章
intuition about the available tools (not necessarily the skills to imple ment all tools in computer code) helps researchers to seek the most appropriate analysis tool for their problem at hand.
对可用工具的直觉理解 (不一定需要掌握用计算机代码实现所有工具的技能) 有助于研究人员为其当前问题寻找最合适的分析工具。
Foundation models in biomedical image analysis
生物医学影像分析中的基础模型
Foundation models, also referred to as foundational models, are a type of deep learning model that is pre-trained on a large, diverse dataset using SSL. Foundation models can be trained on any data, including images or text or both. The goal of these models is to learn general features and patterns that can be applied to a wide range oftasks,rather than being trained for a specific task from scratch.Forexample,various microscopy images, including bright field and fluorescent imaging, obtained under different conditions and with different microscopes, are used to train a foundation model. These models learn general features and patterns across the data they are trained on; for example, a model trained on microscopy images learns about intensities, shapes and structure relations.Sucha pre-training allows the model to capture a broad understanding of the data, which can then be fine-tuned for specific tasks using smaller labelled datasets. Since the early 2020s, foundation models have emerged in biomedical image analysis, enabled by the advent of SSL9°. Foundation models are typically used as a starting point to be fine-tuned using a smaller, labelled dataset, yielding more focused, specialized models. In medical image analysis, this procedure can yield better models for any task, for example, image classification tasks8.18.97. Similar foundation models are being developed for other medical imaging modalities, including cross-modality applications(Fig.1d).Cross-modality applications refer to A I systems that can process and integrate information from different data types, such as combining radiology images with clinical notes or pathology images with genomic data98, or generally, images and text97. To do so requires substantial computational power and engineering resources, as wellas access to extensive datasets.Ideally,acomputing cluster with multiple,oreven dozens of, graphics processing units (GPUs) is necessary. In 2023, various research groups and commercial entities released open-source foundation models, making them widely access ble 9799-101.
基础模型 (Foundation Model) ,也称为基础性模型,是一种通过自监督学习在大型多样化数据集上预训练的深度学习模型。基础模型可以训练任何数据,包括图像、文本或两者兼有。这些模型的目标是学习可广泛应用于各种任务的通用特征和模式,而非针对特定任务从头训练。例如,使用在不同条件下通过不同显微镜获取的各类显微图像(包括明场和荧光成像)来训练基础模型。这些模型能够从训练数据中学习通用特征和模式;例如,在显微图像上训练的模型会学习强度、形状和结构关系。这种预训练使模型能够掌握对数据的广泛理解,随后可通过较小的标注数据集针对具体任务进行微调。
自2020年代初以来,随着自监督学习的出现,基础模型开始在生物医学图像分析领域涌现。基础模型通常作为起点,通过使用较小的标注数据集进行微调,从而产生更专注的专业化模型。在医学图像分析中,这种方法可为任何任务(例如图像分类任务)生成更优的模型。类似的基模型正在被开发用于其他医学成像模态,包括跨模态应用(图1d)。跨模态应用指能够处理和整合不同数据类型信息的AI系统,例如将放射学图像与临床记录结合,或将病理图像与基因组数据整合,或通常意义上的图像与文本结合。
实现这些应用需要大量计算能力、工程资源以及获取广泛数据集的权限。理想情况下,需要配备多个甚至数十个图形处理器 (GPU) 的计算集群。2023年,多个研究团队和商业实体发布了开源基础模型,使其得以广泛使用。
In summary,the rise of foundation models marks an advancement in cancer research9, which has already led to practical improve ments in performance in relevant tasks, enabling training of more effective models with less data97101,102(Fig. 2a,b). Utilizing these models still requires Python programming skills, but future developments may com mod it ize this technology further, making it accessible to researchers with limited or no programming expertise.
总之, 基础模型 (Foundation Model) 的兴起标志着癌症研究的重大进展 [9], 这些模型已在相关任务中带来实际性能提升, 能够用更少的数据训练出更有效的模型 [97,101,102] (图 2a,b) 。使用这些模型目前仍需 Python语言 编程技能, 但未来的发展可能进一步普及这项技术, 使编程经验有限或完全不懂编程的研究人员也能使用。
Al for language Natural language processing
面向语言的自然语言处理
Natural language is unstructured data, of which computer-based analysis has been highly challenging. Since the early 2020s, the advent of large language models (LLMs) has markedly improved the capability of computer-based methodsfor NLP,and are nowthe state-of-the-art approach for processing any text103(Fig.3a). NLPis abroad field within AIthat focuses onthe interaction between computers and human language, whereas LLMs are a specific type of deep learning model used for NLP tasks. LLMs, which are based on transformer architectures, havebecome the most popularand effective method for NLPin recent years. LLMs are part of generative Al, which means they can not only rephrase,summarize or translate text but also synthesize new text. The increasing availability and standardization of LLMs have made NLP methods more accessible, enabling non-experts to solve NLP tasks with off-the-shelf models, such as OpenAl's chatGPT. Beyond storing and retrieving knowledge, LLMs can reason about text104.15s, are capable of translating and stylistically altering text,and can extract structured information from radiology reports106,pathology reports107 and medical notes10°. LLMs also have medical knowledge 109, and can use this, for example, to give medical recommendations based on imaging reportsllo. These capabilities are particularly impactful in cancer research, which relies on text on several levels - from logging initial ideas and experimental data to exchanging insights and disseminating scientific findings (Fig. 3b). The increasing availability and adoption of LLMs in cancer research settings is expected to substantially impact research methodologies!",although their clinical application is still in the early stages due to regulatory and validation challenges 2.
自然语言是非结构化数据,其基于计算机的分析一直极具挑战性。自2020年代初以来,大语言模型 (LLM) 的出现显著提升了基于计算机的自然语言处理方法的能力,并已成为处理任何文本的最先进方法[103] (图 3a)。自然语言处理是人工智能中的一个广泛领域,专注于计算机与人类语言之间的交互,而大语言模型是用于自然语言处理任务的一种特定类型的深度学习模型。基于Transformer架构的大语言模型,已成为近年来最流行和有效的自然语言处理方法。大语言模型是生成式AI (Generative AI) 的一部分,这意味着它们不仅可以重述、总结或翻译文本,还能合成新文本。大语言模型日益增长的可用性和标准化使得自然语言处理方法更易于使用,让非专家能够使用现成模型解决自然语言处理任务,例如OpenAI的ChatGPT。除了存储和检索知识外,大语言模型还能对文本进行推理[104,105],能够翻译和风格化修改文本,并能从放射学报告[106]、病理学报告[107]和医疗记录[108]中提取结构化信息。大语言模型还具备医学知识[109],并能运用这些知识,例如根据影像报告提供医疗建议[110]。这些能力在癌症研究中尤其具有影响力,因为癌症研究在多个层面依赖文本——从记录初步想法和实验数据到交流见解和传播科学发现 (图 3b)。大语言模型在癌症研究环境中日益增长的可用性和采用预计将显著影响研究方法,尽管由于监管和验证方面的挑战,其临床应用仍处于早期阶段[2]。
The advent of LLMs opens a myriad of applications in cancer research, many of which are only beginning to be explored 103. Researchers can now use these readily available modelsfor various tasks:parsing and refining their ideas, summarizing laboratory notebooks, reasoning about complex concepts, acquiring new skills, condensing docu ments and disseminating research findings more effectively 103,1o5,1,14. As NLP evolves further, we are continuously uncovering new uses,each with the potential to enhance the efficiency and scope of academic research and, ultimately, change the clinical practice of oncology!1s. Of note, LLMs signify a departure from traditional machine learning approaches, as classical machine learning requires specific datasets for problem-solving, followed by model training on these data and evaluation. LLMs such as OpenAl's popular 'generative pre-trained transformer(GPT)'models,including the models underlying chat GP T, are by design foundation models9°. Trained on extensive text corpora covering various domains using SSLli6, including on medical and scientific text, LLMs accumulate a broad knowledge base. Consequently, researchers generally donot retrain these modelsfrom scratch dueto the immense computational resources required. Instead, foundation LLMs are typically applied directly to research tasks in a zero-shot way.
大语言模型 (LLM) 的出现为癌症研究开辟了无数应用领域,其中许多才刚刚开始被探索[103]。研究人员现在可以利用这些现成模型完成多种任务:梳理和完善想法、总结实验笔记、进行复杂概念推理、获取新技能、精简文档以及更有效地传播研究成果[103,105,1,14]。随着自然语言处理 (NLP) 的进一步发展,我们不断发现新的应用场景,每个场景都有可能提升学术研究的效率与范围,并最终改变肿瘤学的临床实践[15]。值得注意的是,大语言模型标志着与传统机器学习方法的分离——经典机器学习需要针对特定问题准备专用数据集,随后基于这些数据进行模型训练和评估。而诸如OpenAI广受欢迎的生成式预训练Transformer (GPT) 模型(包括ChatGPT底层模型)本质上都是基础模型[90]。通过自监督学习 (SSL) [116] 在涵盖多个领域的海量文本语料(包括医学和科学文本)上进行训练,大语言模型积累了广泛的知识基础。因此,由于所需计算资源巨大,研究人员通常不会从头开始重新训练这些模型,而是以零样本 (Zero-shot) 方式直接将基础大语言模型应用于研究任务。
Zero-shot applications. The prevalent method is called zero-shot application,in whichan LLMisused directly on a task without previous specific training on any training examples 104. For instance, a researcher can prompt the model to structure or summarize their unstructured notes by inputting the text into an LLM, which processes the text and returns a summary. The efficacy of zero-shot applications relies on prompt engineering ll 7. Effectively phrasing prompts can markedly enhance outcomes without the need for additional data or algorithmic adjustments. In medical applications, zero-shot methods have been used to summarize unstructured radiology reports106,pathology reports107 or clinical notesi08,1s furthermore,to provide treatment recommendations based on radiology images or to respond to medical examination questions 109. Zero-shot applications often work well on simple tasks, and whenever this is the case, it is arguably the easiest and fastest way of using anLLM.
零样本应用。主流方法称为零样本应用,即直接使用大语言模型处理任务,无需事先基于任何训练样本进行特定训练 [104]。例如,研究人员可通过将文本输入大语言模型,提示其整理或总结非结构化笔记,模型处理文本后返回摘要。零样本应用的效能依赖于提示工程 [11,17]。有效组织提示词可在无需额外数据或算法调整的情况下显著提升效果。在医疗应用中,零样本方法已被用于总结非结构化放射学报告 [106]、病理学报告 [107] 或临床笔记 [108,115],还可根据放射影像提供治疗建议或回答医学考试问题 [109]。零样本应用通常在简单任务上表现良好,若适用该方法,无疑是使用大语言模型最简便快捷的途径。
Beyond zero-shot applications. When zero-shot applications are insufficient,researchers may resort to few-shot learning, which involves providing the LLM with a handful of examples to demonstrate the desired task. Few-shot learning can be achieved without retraining the modelat all. The most convenient way to approach few-shot learning with LLMs is with in-context learning. Here, examples are passed to the model as part of the promptl9. For example, a researcher using an LLM to format scientific citations in a specific style could prompt the model with a few examples of such citations. The LLM would then be able to reformat new citations, using the knowledge from the examples
超越零样本应用。当零样本应用不够用时,研究人员可能会采用少样本学习,即为大语言模型提供少量示例来展示所需任务。少样本学习完全无需重新训练模型即可实现。最便捷的大语言模型少样本学习方法是上下文学习。此时,示例作为提示的一部分传递给模型[9]。例如,研究人员若想使用大语言模型按照特定格式整理科学引文,可以向模型输入几个该格式的引文示例。大语言模型随后就能利用这些示例中的知识来重新格式化新的引文。
Review article
综述文章


Fig.3 | Text-based hypothetical AI workflows in cancer research. a, The evolution of natural language processing (NLP) models has seena shift from recurrent neural network(RNN)architectures to transformer architectures. In these transformer models,input tokens (forexample,words)arefirst converted into vectors through a crucial part of the NLP model:the embedding layer.Here, words or phrases are transformed into numerical representations,enabling the model to process language.These vectors,along with position and segment embeddings(which add contextual information about the order and relationship oftokens),are then passed through'transformer encoders'Transformer encoders are advanced components of the model that process the input by considering the context ofeach word in relation to the others,enabling the modelto understand complex language patterns. The output layer ofthe transformer model generates the final predictions or representations based on the processed input, which can be used for various downstream tasks such as text classification,named entity recognition or text generation.b, Trained NLP models can be applied to various tasks in cancer research.For example,they can convert unstructured clinical notes or research papers into structured formats, makingthe information more easily accessible and analysable. This can be achieved by training the modelona large corpus of annotated text data, in which the desired structured format is provided aslabels.The model learns to identify key information and map it to the appropriate fields in the structured format. NLP models can also summarize lengthy documents,such asresearch articles or clinical trial reports, highlighting the key points and findings. c, The training and fine-tuning ofthese models depend on the dataset size and computational power. De novo training of large language models (LLMs) requires internet-scale datasets,containing billions of words, and massive computing infrastructure.However, once these large models are pre-trained, they can be fine-tuned for specific tasks using much smaller domain-specific datasets,often in the range of afew hundred to afew thousand examples. This fine-tuning process is less computationally demanding and can be performed on smaller-scale infrastructure.For certain tasks,LL Ms can be used in a'zero-shot manner, in which they perform thetask without any additional training,relying solely on their pre-existing knowledge.Thiszero-shot capability allows LL Ms to be applied to a wide range of tasks,even on resource-constrained devices such as mobile phones.In-context learning and retrieval-augmented generation (RAG)are alternative approaches,in whichthe modelis provided withafew examples of the desired task as part ofthe input prompt orby providing the model with additional documents,respectively,enablingit to adapt to the task without explicit fine-tuning. These various approaches have made language understanding and generation capabilities more accessible and efficient.
图 3 | 癌症研究中基于文本的假设性 AI 工作流程。a, 自然语言处理 (NLP) 模型的演变经历了从循环神经网络 (RNN) 架构向 Transformer 架构的转变。在这些 Transformer 模型中,输入 Token (例如单词) 首先通过 NLP 模型的关键部分——嵌入层转换为向量。在此过程中,单词或短语被转化为数值表示,使模型能够处理语言。这些向量与位置嵌入和分段嵌入 (用于添加关于 Token 顺序和关系的上下文信息) 一起被传递至 "Transformer 编码器"。Transformer 编码器是模型的高级组件,通过考虑每个单词与其他单词的上下文关系来处理输入,使模型能够理解复杂的语言模式。Transformer 模型的输出层基于处理后的输入生成最终预测或表示,可用于各种下游任务,如文本分类、命名实体识别或文本生成。b, 经过训练的 NLP 模型可应用于癌症研究中的各种任务。例如,它们可以将非结构化的临床笔记或研究论文转换为结构化格式,使信息更易于访问和分析。这可以通过在大量带标注的文本数据上训练模型来实现,其中所需的结构化格式作为标签提供。模型学会识别关键信息并将其映射到结构化格式的相应字段中。NLP 模型还能总结冗长文档 (如研究文章或临床试验报告),突出关键点和发现。c, 这些模型的训练和微调取决于数据集大小和计算能力。大语言模型 (LLM) 的从头训练需要互联网规模的数据集 (包含数十亿单词) 和大规模计算基础设施。然而,一旦这些大型模型经过预训练,就可以使用小得多的领域特定数据集 (通常为数百到数千个示例) 针对特定任务进行微调。这种微调过程对计算资源的要求较低,可在较小规模的基础设施上执行。对于某些任务,大语言模型可以 "零样本" 方式使用,即无需任何额外训练,仅依靠其已有知识执行任务。这种零样本能力使大语言模型可应用于广泛任务,甚至在手机等资源受限设备上也能运行。上下文学习 (in-context learning) 和检索增强生成 (RAG) 是两种替代方法:前者在输入提示中提供少量任务示例,后者则为模型提供额外文档,使模型无需显式微调即可适应任务。这些多样化方法使语言理解和生成能力变得更易获取且更高效。
in the prompt, without ever having been retrained in the classical way. In-context learning is constrained by the maximum input size of models, which for many models does not exceed tens of pages oftext.
在提示中,无需以传统方式进行重新训练。上下文学习受到模型最大输入长度的限制,许多模型的输入上限通常不超过几十页文本。
Another common approach for providing a model with context is retrieval-augmented generation (RAG). In RAG, an LLM can embed documents in a vector representation and can later access this information when promptedi2°. RAG is not limited by document size and is conveniently available through programming interfaces or user-facing online tools.RAG can be used,for example,to provide documents with hundreds of pages of text to an LLM and task it to answer questions based on the knowledge in the documents.
另一种为模型提供上下文的常见方法是检索增强生成 (RAG) 。在 RAG 中,大语言模型可以将文档嵌入向量表示中,并在后续被提示时访问这些信息 [20] 。RAG 不受文档大小限制,且可通过编程接口或面向用户的在线工具便捷使用。例如,RAG 可用于将数百页文本的文档提供给大语言模型,并让其根据文档中的知识回答问题。
Finally,ifthese approaches do not succeed because the LL M does not have enough specialized knowledge, the last resort is to fine-tune the LLM with a custom dataset.Forexample,the medicine-specific LLM NYUTron was specifically trained on patient electronic health records (EHRs),enabling it to predict clinical outcomes 1.Datasetsto train LLMs from scratch, or to fine-tune existing LLMs, need to be substantial in size and require substantial computational resources. Commercially available LLMs such as OpenAl's GPT-4 or Anthropic's Claude 3 were probably trained on the entirety of publicly available data on the internet.
最后,如果这些方法因大语言模型 (LLM) 缺乏足够专业知识而未能成功,最终手段是使用定制数据集对大语言模型进行微调。例如,医学专用大语言模型 NYUTron 专门基于患者电子健康记录 (EHRs) 进行训练,使其能够预测临床结局 [1]。用于从头训练大语言模型或微调现有大语言模型的数据集需要具备足够规模,并需要大量计算资源。诸如 OpenAI 的 GPT-4 或 Anthropic 的 Claude 3 等商用大语言模型,很可能是在互联网全部公开数据上进行训练的。
NYUTron was trained on routine clinical notes from approximately 750,0o0 patients l i. Because datasets at this scale and the hardware required to train LLMs on datasets this large are not available to most researchers, zero-shot application and few-shot learning are typically the go-to approaches. In practice,a hybrid approach combining different optimization strategies is often used, involving iterative cycles of prompt engineering,few-shot learning, RAG implementation and ultimately fine-tuning to measure improvements inthe performance of the $\mathbf{LLM}^{122,123}$ . In summary,the most substantial contribution of LLMs to cancer research might not bethe human-like text processing capabilities per se,butthe complete upending of traditional machine learning workflows from rigid train-test-evaluate cycles to zero-shot applications (Fig.3c). In addition,the recent advancements in LLMs will conceivably lead to the advent of multimodal, multi-purpose Al models in the future° (Box 3).
NYUTron 基于约 75 万名患者的常规临床笔记进行训练 [1]。由于这种规模的数据集以及在此类大型数据集上训练大语言模型所需的硬件对大多数研究者而言难以获取,零样本应用和少样本学习通常成为首选方案。实践中常采用结合多种优化策略的混合方法,包括迭代式的提示工程、少样本学习、RAG 实现,最终通过微调来评估大语言模型性能的提升效果 [122, 123]。总体而言,大语言模型对癌症研究最重大的贡献或许并非类人的文本处理能力本身,而是彻底颠覆了传统机器学习工作流程——从僵化的训练-测试-评估循环转向零样本应用 (图 3c)。此外,大语言模型的最新进展预计将推动未来多模态通用人工智能模型的出现 (框 3)。
How to use large language models
如何使用大语言模型
LLMs typically run on a server in a data centre and users can access them through web interfaces, mobile applications or access them via an application programming interface (API). For example, the LLMs GPT-4 (developed by OpenAl), Gemini1.5 (developed by Google) and
大型语言模型 (LLM) 通常运行在数据中心的服务器上,用户可以通过网页界面、移动应用程序或应用程序编程接口 (API) 进行访问。例如,由 OpenAI 开发的 GPT-4、由 Google 开发的 Gemini 1.5 等大型语言模型。
Review article
综述文章
Box3
From multimodal LLMs to generalist medical Al
从多模态大语言模型到通用医学人工智能
Large language models (LLMs), in their simplest form, are just for text. However, since 2022, LLMs have evolved substantially, resulting in multimodal LLMs that integrate text and image processing. These models can be designed as singular entities with image-perception and text-perception capabilities, or as distinct image processing and text encoding parts amalgamated together. For example, the current version of the popular LLM chatGPT accepts images as well as text as an input. This multimodal approach blurs the lines between natural language processing and computer vision, offering superior pattern recognition and analytical skills96.171. In cancer research, these advancements are particularly promising. Multimodal models cannow analyse complex his to pathology images and engage in conversational interactions 135. Ultimately, such models can be trained on diverse data types and offer a unified solution for diverse tasks, such as interpreting microscopy and X-ray images, and processing clinical notes. However, although the technology exists and is rapidly advancing, several practical and regulatory challenges remain172. On the basis of these multimodal LLMs, zero-shot and few-shot learning approaches can be used to process images.
大语言模型 (LLM) 在最简单的形式下仅处理文本。但自2022年以来,大语言模型已取得长足发展,催生出能同时处理文本与图像的多模态大语言模型。这类模型可设计为兼具图像感知与文本感知能力的单一实体,也可由独立的图像处理模块和文本编码模块组合而成。例如当前流行的ChatGPT大语言模型就能同时接收图像和文本输入。这种多模态方法模糊了自然语言处理与计算机视觉的界限,提供了卓越的模式识别与分析能力[96][171]。在癌症研究领域,这些进展尤其令人振奋。多模态模型现已能分析复杂的组织病理学图像并进行对话交互[135]。最终,此类模型可通过多种数据类型进行训练,为显微成像、X射线图像解读及临床笔记处理等多样化任务提供统一解决方案。然而尽管技术已然存在并快速发展,仍存在诸多实践与监管层面的挑战[172]。基于这些多模态大语言模型,零样本和少样本学习方法可被应用于图像处理任务。
The ultimate goal of deep learning is to develop a single multimodal, multitask model capable of simultaneously addressing a wide range of tasks. Such a model would be truly'generalist'96. For example, instead of having three separate models for radiology image analysis, pathology image analysis and genomic data analysis, a single model could integrate the information from all these modalities and yield a single prediction. This would reduce the number of models that are needed in research workflows and increase their performance by exploiting synergies between data types. In the past few years, initial proof-of-concept studies have shown that such synergies exist between genomic data and histology 48.173,radiology and histology 130, radiology and clinical data83 and between different his to logical stains129. These recent studies are just the start of a broader trend from single-purpose, single-modal models towards multi-purpose, multimodal models.
深度学习的终极目标是开发出能够同时处理广泛任务的多模态、多任务单一模型。这样的模型将成为真正的"通用型"模型。例如,无需为放射影像分析、病理图像分析和基因组数据分析分别建立三个独立模型,单一模型就能整合所有模态的信息并给出统一预测。这将减少研究流程中所需的模型数量,并通过利用数据类型间的协同效应来提升性能。过去几年中,初步概念验证研究已证实基因组数据与组织学、放射学与组织学、放射学与临床数据以及不同组织学染色之间均存在此类协同效应。这些最新研究仅仅是从单一用途、单一模态模型向多用途、多模态模型广泛转型的开端。
Extending this, generalist deep learning models could enable an LLM-style user interaction to make model predictions more explain able. By incorporating NLP capabilities, generalist models could provide intuitive explanations for their predictions, similar to how LLMs can generate human-readable text. More specifically, the relatively new concept of generalist medical Al (GMAl) holds the potential to impact cancer research in the coming years96. The proficiency of GMAl in processing diverse data types, from imaging to genetic information, could markedly enhance diagnostic and prognostic methods in oncology. The shift from task-specific models to a more holistic and integrated approach could mark a substantial transformation in the application of Al in cancer research and oncology.
在此基础上,通用深度学习模型可实现类似大语言模型的用户交互模式,从而增强模型预测的可解释性。通过整合自然语言处理能力,通用模型能够为预测结果提供直观解释,类似于大语言模型生成人类可读文本的机制。更具体地说,相对新兴的通用医疗人工智能概念有望在未来几年影响癌症研究[96]。通用医疗人工智能在处理从影像到遗传信息等多元数据类型方面的熟练度,可显著提升肿瘤学的诊断与预后方法。从任务专用模型转向更整体化、集成化的方法,可能标志着人工智能在癌症研究与肿瘤学应用领域的重大变革。
Claude 3 (developed by Anthropic) are highly advanced LLMs and are typically accessed through web interfaces 104,124. Increasingly, LLMs are permeating many aspects of our daily life. They are embedded in more specialized software such as in office software to create presentation slides, ine-mail software for tasks such as drafting responses, prioritizing incoming messages or summarizing e-mail threads, or in task management software for generating task descriptions, setting reminders or providing suggestions for task prior it iz ation. In the future, LLMs could be embedded in many types of scientific software, for example, software for laboratory notebook management or many other types of software to manage the daily workflows in research.
Claude 3 (由 Anthropic 开发) 是高度先进的大语言模型 (LLM) , 通常通过网络界面进行访问 [104,124]. 日益普及的大语言模型正渗透到我们日常生活的诸多方面. 它们被嵌入到更专业的软件中, 例如办公软件中用于创建演示文稿幻灯片, 电子邮件软件中用于起草回复, 对接收消息进行优先级排序或总结电子邮件线程, 或在任务管理软件中用于生成任务描述, 设置提醒或为任务优先级排序提供建议. 未来, 大语言模型可能会嵌入到多种类型的科学软件中, 例如实验室记录管理软件或许多其他类型用于管理研究日常工作流程的软件.
There are multiple ways to interact with LLMs. First, many stateof-the-art models such as GP T-4,Gemini1.5 or Claude 3 are owned and operated by technology corporations and can be accessed through online browser interfaces or mobile phone applications provided by these companies. This option is appropriate for those who prefer a direct and easy-to-use interface.Second,itis possibleto utilize theAPI, which allows for more flexibility and integration into specific projects or workflows. Third, there is the option of using LLM-enabled softwares, such as you.com. These platforms integrate an LLM with additional fun ction ali ties such as integrated search capabilities and data visualization and analysis tools, providing a more comprehensive toolset for users seeking an enhanced experience with advanced features25. However, not all LLMs runon the servers ofbig corporations. There is an increasingly active open-source ecosystem of LLMs, including Llamal2°, Mixtral and many of their derivatives which are,forexample,available on the Hugging Face web portal. Here, many open-source LLMs are available that can be run on standard commercial hardware and used locally for biomedical text processing lo 8. This allows researchers to maintain control over their data and ensures confidentiality, as the data do not need to be uploaded to external servers.
与大语言模型 (LLM) 交互存在多种方式。首先,许多前沿模型(如 GPT-4、Gemini 1.5 或 Claude 3)由科技公司所有并运营,可通过这些公司提供的在线浏览器界面或手机应用程序进行访问。该选项适合偏好直接易用界面的用户。其次,可通过应用程序编程接口 (API) 进行调用,这为特定项目或工作流程提供了更高灵活性和集成能力。第三,可使用支持大语言模型的软件平台(如 you.com),这些平台将大语言模型与集成搜索功能、数据可视化分析工具等附加功能相结合,为追求增强体验的用户提供更全面的工具集 [25]。
然而并非所有大语言模型都运行于大型企业的服务器。目前日益活跃的开源大语言模型生态体系包含 Llama 2、Mixtral 及其众多衍生模型(例如可通过 Hugging Face 网络平台获取)。这些开源大语言模型可在标准商用硬件上本地运行,用于生物医学文本处理 [10][18]。这使得研究人员能够保持对数据的控制权并确保机密性,因为数据无需上传至外部服务器。
Emerging uses of Al Multimodal Al
人工智能的新兴应用:多模态人工智能
Multimodal data fusion. An emerging and fundamental improvement for deep learning lies in the ability to effectively handle and integrate information from diverse modalities 127. For example, in cancer histopathology, some visual patterns can only be interpreted by humans or computers in the context of additional information.With current technical capabilities, it is now feasible to combine multiple data types as input for a single deep learning system. In cancer research and clinical diagnostics,it seems intuitive that enriching image data with additional context information can improve classifications or predictions, as images are usually interpreted in the context of additional data. Recent studies have incorporated small amounts oftabular data,such as the diagnosis, age and sex of the patient, alongside image data to improve the performance of deep learning models in various cancerrelated tasks, for example, in his to pathology image process inglis Similarly,in radiology image processing, integrating clinical data with images has been shown to enhance the accuracy of deep learningbased diagnoses 83. In addition, using H&E-stained slides together with immuno his to chemistry-stained slides as ajoint input into a single deep learning system can better predict individual patient risks129. Furthermore, deep learning systems can integrate genomic data with routine
多模态数据融合。深度学习一个新兴且根本性的改进在于能够有效处理和整合来自不同模态的信息 127。例如,在癌症组织病理学中,某些视觉模式只有在附加信息的背景下才能被人类或计算机解读。凭借现有技术能力,将多种数据类型作为单一深度学习系统的输入已成为可能。在癌症研究和临床诊断中,用额外背景信息丰富图像数据可以改进分类或预测,这似乎是直观的,因为图像通常是在附加数据的背景下解读的。最近的研究纳入了少量表格数据,例如患者的诊断、年龄和性别,以及图像数据,以提高深度学习模型在各种癌症相关任务中的性能,例如在组织病理学图像处理中。类似地,在放射学图像处理中,将临床数据与图像整合已被证明可以提高基于深度学习的诊断的准确性 83。此外,将H&E染色切片与免疫组织化学染色切片一起作为单一深度学习系统的联合输入,可以更好地预测个体患者风险 129。此外,深度学习系统可以将基因组数据与常规
Review article
综述文章
pathology 48,which is particularly useful as together this provides the spatial resolution, and molecular details. Several studies have shown such a synergy between spatial (images) and non-spatial (molecular) data can be exploited by multimodal deep learning systems48,127,130.131. For non-computational cancer researchers entering the field of multimodal deep learning,it is crucial to understand that data availability is a primary challenge. Obtaining multimodal data for more than a few hundred patients is often extremely difficult in practice. Therefore, the key lesson is that all available anonymized data with common patient identifiers should be stored securely in an appropriate repository for potential use in future multimodal deep learning exploratory studies.
病理学 48, 这种结合特别有价值, 因为它能同时提供空间分辨率和分子细节。多项研究表明, 多模态深度学习系统可以利用空间 (图像) 和非空间 (分子) 数据之间的这种协同效应 48,127,130.131。对于刚进入多模态深度学习领域的非计算癌症研究人员而言, 必须认识到数据可获得性是首要挑战。在实践中, 获取超过数百名患者的多模态数据通常极其困难。因此, 关键经验是所有带有通用患者标识符的可用匿名数据都应安全存储在适当的存储库中, 以备未来多模态深度学习探索性研究使用。
Vision LLMs.Vision LLMs differfrom multimodal data fusion in that they arepre-trained on large,diverse datasets containing both text and images, allowing them to process and generate both modalities simultaneously. Bycontrast,multimodal data fusion typically involves combining d ifferent data types (for example, images, text and genomic data) using task-specific deep learning models that are trained from scratch or finetuned from pre-trained single-modality models. Vision LLMsoffer a more general and flexible approach to multimodal learning, as they can be appliedto a wide range oftasks with minimal fine-tuning, whereas multimodal data fusion models are usually designed and optimized for specific applications.LLMs with vision capabilities,such as Claude 3 orG PT-4,can process image data alongside text.Some applications of such vision LLMs have already been explored incancer research 132.Open-source LLMs,such as Llava133 and its medical specialist model Llava-Medl34, demonstrate promising capabilities in biomedical image analysis. Another example is Path Chat 135, a chatbot designed for pathologists. PathChat is built on topofa vision LLM architecture and canengage in interactive discussions with pathologists, providing insights and answering questions related to specific specialized use cases.Most ofthese technical advancements occurred in 2023 and 2024, indicating a dynamic future for Alin cancer research.As vision LLMs continue to evolve and become more accessible, they are likely to have an increasingly important role in integrating multiple data modalities and supporting various aspects of cancer research and ultimately for clinical practice in oncology.
视觉大语言模型 (Vision LLMs) 。视觉大语言模型与多模态数据融合的区别在于,它们是在包含文本和图像的大型多样化数据集上进行预训练的,使其能够同时处理和生成两种模态。相比之下,多模态数据融合通常涉及使用特定任务的深度学习模型(例如图像、文本和基因组数据)进行组合,这些模型是从头开始训练或从预训练的单模态模型微调而来。视觉大语言模型为多模态学习提供了更通用灵活的方法,因为它们只需最小程度的微调即可应用于广泛任务,而多模态数据融合模型通常针对特定应用进行设计和优化。具备视觉能力的大语言模型(如 Claude 3 或 GPT-4)可同时处理图像和文本数据。此类视觉大语言模型在癌症研究中的部分应用已得到探索 [132]。开源大语言模型(如 Llava [133] 及其医学专用模型 Llava-Med [134])在生物医学图像分析中展现出潜力。另一个案例是面向病理学家设计的聊天机器人 PathChat [135],该程序基于视觉大语言模型架构构建,可与病理学家进行交互式讨论,针对特定专业用例提供见解并解答问题。这些技术突破大多发生于2023至2024年,预示着人工智能在癌症研究领域将迎来充满活力的未来。随着视觉大语言模型的持续演进和普及,它们将在整合多数据模态、支持癌症研究各环节乃至最终应用于肿瘤学临床实践方面发挥日益重要的作用。
Analysing the genome and the proteome
分析基因组与蛋白质组
LLMs excel not only in processing natural language but can also be applied to any biological sequence data.Genomic sequence data represent another language that LLM-like transformer models can interpret. In the field of genomic data analysis, there are established classical bioinformatics pipelines that do not depend on LLMs or deep learning, and are the state of the art to extract scientifically relevant or clinically actionable information from raw sequencing reads136-138. These pipelines work well and are understandable,so there is no pressing need to replace them with deep learning. However, more subtle genomic signatures, such as specific DNA damage signatures, could yet be discovered by deep learning i 39. An argument for using deep learning in genome data analysis is to set the stage for multimodal models48. Beyond parsing genomic sequences, deep learning has been used for many specialized tasks in molecular biology,including the prediction ofthe likelihood of antigen presentation from a gene of interest i 4 o. The development of models that accurately predict the three-dimensional structure of proteins from their genomic sequence using deep learning, termed AlphaFold models141, has solved a decades-old puzzle in biology. This has enabled arange of downstream applications 14 such as missense predictions for all possible human single amino acid changes143 or to analyse the similarity of all known proteins 144.
大语言模型 (LLM) 不仅在自然语言处理方面表现出色,还能应用于任何生物序列数据。基因组序列数据代表了另一种类似 Transformer 的模型可以解读的语言。在基因组数据分析领域,存在不依赖大语言模型或深度学习的经典生物信息学流程,这些流程是从原始测序数据中提取科学相关或临床可操作信息的先进方法 [136-138]。这些流程运行良好且易于理解,因此没有迫切需要将其替换为深度学习。然而,更细微的基因组特征,例如特定的 DNA 损伤特征,仍可能通过深度学习被发现 [139]。在基因组数据分析中使用深度学习的一个论据是为多模态模型奠定基础 [48]。除了解析基因组序列,深度学习已被用于分子生物学中的许多专业任务,包括从目标基因预测抗原呈递可能性 [140]。通过深度学习从基因组序列准确预测蛋白质三维结构的模型开发,被称为 AlphaFold 模型 [141],解决了生物学中一个存在数十年的难题。这使得一系列下游应用成为可能 [142],例如预测所有可能的人类单氨基酸错义突变 [143],或分析所有已知蛋白质的相似性 [144]。
Drug discovery and clinical trial optimization
药物发现与临床试验优化
Machine learning has been broadly used in drug discovery for decades145, but, recently, the success of large transformer models have put Al-based tools in the spotlight ofthe field of drug discovery. Deep learning models such as AlphaFold are highly efficient at predicting protein structures, a potentially useful tool in drug discovery. These models could be extended to predict the binding of potential drug candidates to functional areas of proteins and also navigate the vast combinatorial landscape of proteins, many of which may possess valuable therapeutic properties 146. Other deep learning models could be trained togenerate specific compounds witha desired property,including small molecules 147, but also RNA or proteins 148. Recent studies have shown that deep learning models can identify chemical substructures linked to antibiotic activity, enabling predictions of new antibiotic classes and substances 149,150. This approach has been used to evaluate millions of compounds,identifying those with potential antibiotic properties and low cyto toxicity to human cells. Of note, among the tested compounds, some showed effectiveness against antibioticresistant bacteria. In addition, machine learning methods have been used to model potential synergies across different drugsi51.
机器学习已在药物发现领域广泛应用数十年 [145] , 但近期大型 Transformer 模型取得的成功使基于人工智能 (AI) 的工具成为药物发现领域的焦点。诸如 AlphaFold 之类的深度学习模型能高效预测蛋白质结构, 这是药物发现中潜在的有用工具。这些模型可扩展用于预测候选药物与蛋白质功能区域的结合, 并能探索庞大的蛋白质组合空间, 其中许多可能具有宝贵治疗特性 [146] 。其他深度学习模型可被训练生成具有特定属性的化合物, 包括小分子 [147] , 以及 RNA 或蛋白质 [148] 。最新研究表明, 深度学习模型能识别与抗生素活性相关的化学亚结构, 从而预测新型抗生素类别和物质 [149,150] 。该方法已用于评估数百万种化合物, 识别出具有潜在抗生素特性且对人类细胞毒性较低的化合物。值得注意的是, 在测试化合物中, 有些显示出对抗耐药菌的有效性。此外, 机器学习方法已被用于模拟不同药物间的潜在协同效应 [151] 。
Another approach is to use LLMs to navigate knowledge graphs in pharmaceutical research, thereby enabling researchers to navigate the existing space faster and potentially more efficiently 152. Knowledge graphs are structured representations of information that capture entities,such as compounds,proteins and diseases, and the relationships between them, such as drug-target interactions or disease-gene associations. By using the large amount of curated information stored in these knowledge graphs, LLMs could help researchers to identify potential drug candidates,predict drug-drug interactions or repurpose existing drugs for new indications. Al-based tools are notjust influencing computational research in drug discovery, but a recent study suggested that a LLM can be coupled with a robotic system to perform independent chemical experimentation 153. In this environment, some authors have advocated for caution154-156. The rapid growth of Al-based drug development companies since the early 2020s, with partnerships with large pharmaceutical firms, highlights the commercial potential of these technologies 15715 s. However, many claims about the capabilities of Al in drug design originate from these companies, introducing potential bias. The true effect of Al-based tools on shortening the path from initial drug conception to clinical trials and ultimate drug approval remains to be empirically demonstrated.
另一种方法是利用大语言模型在药物研究中导航知识图谱,从而使研究人员能够更快且可能更高效地探索现有空间[152]。知识图谱是结构化的信息表示方式,可捕获化合物、蛋白质和疾病等实体及其相互关系(如药物-靶点相互作用或疾病-基因关联)。通过利用这些知识图谱中存储的大量经过整理的信息,大语言模型可以帮助研究人员识别潜在候选药物、预测药物相互作用或将现有药物重新用于新适应症。基于人工智能的工具不仅正在影响药物发现领域的计算研究,最近一项研究表明,大语言模型可以与机器人系统结合以进行独立的化学实验[153]。在此环境下,部分作者主张保持谨慎[154-156]。自2020年代初以来,基于人工智能的药物开发公司通过与大型制药企业建立合作伙伴关系实现快速增长,凸显了这些技术的商业潜力[157,158]。然而,关于AI在药物设计能力的许多主张都源自这些公司,可能存在潜在偏见。基于AI的工具在缩短从初始药物构想到临床试验及最终药物获批路径方面的实际效果,仍有待实证验证。
After a candidate drug has been developed, it needs to proceed to clinical trials, and currently $90%$ of these fail to reach clinical implement ation 159. Al-based tools hold the potential to improve this at many levels16o. For example, Al could optimize trial recruitment by refining eligibility criteria and improving the efficiency of candidate identification,screening and inclusion.Inaddition, LLMs could parse EHR data and automatically flag candidates for a specific triall61. Similarly, deeplearning modelsthat predict genetic alterations from pathology slides could help to identify candidate patients for a trial faster17. Few studies have empirically tested this,presumably because the enabling technologies arestill new;the contemporaneous LL MG PT-4 was only released in 2023. Furthermore, Al application to medical imaging could aid in identifying early signs of impending progression, assisting in treatment decisions better than subjective description of the changes in tumour size in colorectal cancer and lung cancer162,163. To implement and validate these emerging Al applications ingenomics and drug discovery,independent scientific evaluation and rigorous clinical trials are essential. It would
候选药物研发成功后需要进入临床试验阶段,目前这些试验中有 $90%$ 未能实现临床应用 [159]。基于人工智能 (AI) 的工具在多个层面具有改善这一现状的潜力 [160]。例如,AI 可通过优化入选标准、提升候选者识别效率、改进筛查与纳入流程来优化试验招募。此外,大语言模型能够解析电子健康记录 (EHR) 数据并自动标记特定试验的候选者 [161]。同样地,基于病理切片预测基因突变的深度学习模型有助于更快识别适合临床试验的候选患者 [17]。目前对此开展实证研究的工作较少,可能因为相关支撑技术仍处于新兴阶段——同期发布的 GPT-4 大语言模型直至 2023 年才问世。此外,AI 在医学影像中的应用有助于识别病情进展的早期征兆,相较于对结直肠癌和肺癌肿瘤大小变化的主观描述,能更有效地辅助治疗决策 [162, 163]。为在基因组学与药物研发领域实施并验证这些新兴 AI 应用,独立的科学评估与严格的临床试验至关重要。
Review article
综述文章
Glossary
术语表
Application programming interface
应用程序编程接口
(API). A set of tools and protocols for building software and applications, enabling software to communicate with Al models.
(API) 。一套用于构建软件和应用程序的工具和协议,使软件能够与AI模型进行通信。
Artificial neural networks
人工神经网络
(ANNs). Computational models loosely inspired by the structure and function of the human brain, consisting of interconnected layers of nodes, called neurons, that process input data and learn to recognize patterns and make decisions.
人工神经网络 (ANNs) 。一种计算模型, 其设计灵感大致来源于人脑的结构和功能, 由相互连接的节点层 (称为神经元) 组成, 这些节点处理输入数据并学习识别模式及做出决策。
Deep learning
深度学习
Deep learning is a subfield of machine learning that uses artificial neural networks with multiple layers, called deep neural networks, to learn and extract highly complex features and patterns from raw input data.
深度学习是机器学习的一个子领域,它使用具有多个层的人工神经网络(称为深度神经网络)来学习和从原始输入数据中提取高度复杂的特征和模式。
Digital images
数字图像
Computational pathology
计算病理学
Visual representations captured and stored in a digital format, consisting of a grid of pixels, with each pixel representing a colour intensity value.
以数字格式捕获和存储的视觉表示,由像素网格组成,每个像素代表颜色强度值。
The use of algorithms, machine learning and image analysis techniques to extract information from digital pathology images.
利用算法、机器学习与图像分析技术从数字病理图像中提取信息。
Digital pathology
数字病理学
Computer vision
计算机视觉
The practice of converting glass slides into digital slides that can be viewed, managed and analysed on a computer.
将玻璃载玻片转换为可在计算机上查看、管理和分析的数字化载玻片的实践。
A field of Al that focuses on enabling computers to analyse and interpret visual data, such as images and videos.
计算机视觉 (Computer Vision) 是人工智能的一个领域,专注于使计算机能够分析和解释图像、视频等视觉数据。
Explain ability methods
可解释性方法
Techniques in Al that provide insights and explanations on how the Al model arrived at its conclusions, thus making the decision-making process of the Al more transparent.
人工智能技术,可提供关于人工智能模型如何得出结论的见解和解释,从而使人工智能的决策过程更加透明。
Convolutional neural networks
卷积神经网络
(CNNs). A type of deep neural network that is especially effective for analysing visual imagery and used in image analysis.
(CNNs). 一种深度神经网络, 特别适用于分析视觉图像并用于图像分析.
Generative Al
生成式AI (Generative AI)
Al systems that can generate new content (text, images or music) that is similar to the content on which it was trained, often creating novel and coherent outputs.
能够生成与训练数据相似的新内容 (文本、图像或音乐) 的人工智能系统,通常能产生新颖且连贯的输出内容。
be beneficial if Al models for drug discovery were open source so that independent researchers could test and improve their performance.
如果用于药物发现的AI模型能够开源,将使独立研究人员能够测试并改进其性能,这将是有益的。
Conclusion
结论
A key challenge in applying Al to cancer research lies in mining the potential of real-world data (RWD) in oncology, which includes EHRs, medical images, tumour samples and blood tests164. Unlike clinical trial data, which is prospectively collected following well-defined proto cols and structured in easily accessible databases,RWD are often unstructured, heterogeneous and scattered across different systems. EHR systems are a particularly interesting data source for real-world Al-based systems121,165,16. Al-based analysis of routine clinical data also holds promise for personalized medicine. Advancing precision oncology will require the development of a larger number of more accurate biomarkers for predicting treatment response, the identification of new druggable targets in cancer cells and an accelerated drug development platform to simplify matching targets with drugs. Here, Al could uncover hidden patterns in molecular profiling to advance this.Although Al has been extensively utilized to uncover patterns in tumour molecular profiling associated with clinical outcomes,such as
将人工智能应用于癌症研究的一个关键挑战在于挖掘肿瘤学领域真实世界数据 (RWD) 的潜力,这些数据包括电子健康记录 (EHR) 、医学影像、肿瘤样本和血液检测[164]。与按照明确定义方案前瞻性收集并存储于易访问数据库的临床试验数据不同,真实世界数据通常是非结构化、异质性且分散在不同系统中。电子健康记录系统对于基于人工智能的真实世界系统而言是特别有价值的数据源[121,165,166]。基于人工智能的常规临床数据分析也为个性化医疗带来希望。推进精准肿瘤学需要开发更多更准确的生物标志物来预测治疗反应,识别癌细胞中新的可成药靶点,以及建立加速药物研发平台以简化靶点与药物的匹配。在这方面,人工智能可通过揭示分子谱中的隐藏模式来推动这一进程。虽然人工智能已被广泛用于发现与临床结果相关的肿瘤分子谱模式,例如
Gigapixel images
十亿像素图像
Extremely high-resolution digital images consisting of 1 billion pixels, obtained by scanning tissue slides with a slide scanner.
由载玻片扫描仪扫描组织切片获得的包含10亿像素的极高分辨率数字图像。
Natural language processing (NLP). A branch of Al that helps computers to analyse, interpret and respond to human language in a useful way.
自然语言处理 (NLP) 。人工智能的一个分支,旨在帮助计算机以有效方式分析、解释和响应人类语言。
Graphics processing units Prompt engineering
图形处理器提示工程
(GPUs). Specialized hardware used to rapidly process large blocks of data simultaneously, used in computer gaming and Al.
(GPUs). 用于快速并行处理大数据块的专用硬件, 广泛应用于电脑游戏和人工智能领域。
Crafting inputs or questions in a way that guides Al models, particularly LLMs, to provide the most effective and accurate responses.
通过精心设计输入或问题的方式,引导AI模型(特别是大语言模型 (LLM))提供最有效和准确的响应。
Large language models
大语言模型
(LLMs). Advanced Al models trained on vast amounts of text data, capable of analysing generating and manipulating human language, often at the human level74.
(LLMs). 在海量文本数据上训练出的先进人工智能模型, 能够分析、生成和处理人类语言, 其能力通常达到人类水平[20]。
Transformers
Transformers
Types of a neural network model that excel at processing sequences of data, such as sentences in text, by focusing on different parts of the sequence to make predictions 175.
一种擅长处理序列数据 (例如文本中的句子) 的神经网络模型, 它通过关注序列的不同部分进行预测 175.
Long short-term memory (LSTM) networks
长短期记忆 (LSTM) 网络
Voxel
Voxel
A type of neural network particularly good at processing sequences of data (such as time series or language), with a capability to remember information for a certain time.
一种特别擅长处理序列数据 (例如时间序列或语言) 的神经网络类型, 具有在一定时间内记住信息的能力。
The three-dimensional equivalent of a pixel in images, representing a value on a regular grid in three-dimensional space, commonly used in medical imaging such as MRI and CT scans.
体素是图像中像素的三维等效物,代表三维空间规则网格上的一个数值,常用于医学成像 (如 MRI 和 CT 扫描) 中。
Machine learning
机器学习
A subset of Al focusing on the development of algorithms and models that enable computers to learn and improve their performance on a specific task without being explicitly instructed how to achieve this.
人工智能的一个子集,专注于开发算法和模型,使计算机能够在没有明确指导的情况下学习并提高其在特定任务上的性能。
prognosis and therapy response l 67, translating these findings into clinical practice remains a challenge. This is partly due to the complexity of cancer biology,the heterogeneity of patient populations,and theneed for rigorous validation of Al-derived biomarkers and drug targets. To fully realize the potential of Alin advancing precision oncology,several key challenges need tobe addressed. These include the integration of multiple data modalities (for example, imaging, genomics and clinical data),the development of explain able and transparent Al models,and the establishment of standards for data sharing and model validation. In addition, closer collaboration between Al researchers,cancer biologists and clinicians will be essential to ensure that Al-based tools are clinically relevant and aligned with the needs of patients and healthcare providers. The rapid advancements in Al, driven by substantial hardware and algorithm improvements, hint at an exciting future. It is advisable that cancer researchers become familiar with these tools and develop an intuition about the potential and the limitations of contemporaneous Also that they areableto navigate a world in which Al-basedtools are ubiquitously used.
预后与治疗反应 l 67,将这些发现转化为临床实践仍面临挑战。这部分归因于癌症生物学的复杂性、患者群体的异质性,以及需要对人工智能衍生的生物标志物和药物靶点进行严格验证。为充分发挥人工智能在推进精准肿瘤学方面的潜力,需要解决若干关键挑战:包括整合多模态数据(例如影像学、基因组学和临床数据)、开发可解释且透明的人工智能模型,以及建立数据共享与模型验证标准。此外,人工智能研究人员、癌症生物学家与临床医生之间更紧密的合作至关重要,这将确保基于人工智能的工具既符合临床需求,又契合患者与医疗提供者的诉求。在硬件与算法大幅改进的推动下,人工智能的快速发展预示着令人振奋的未来。建议癌症研究者熟悉这些工具,对当代人工智能的潜力与局限形成直觉认知,并适应这个普遍使用人工智能工具的新环境。
Published online: 16 May 2024
在线发布:2024年5月16日
综述文章
Review article
综述文章
Acknowledgements
致谢
R.P.-L.is supported by LaCaixa Foundation,a CRIS Foundation Talent Award (TALENT19-O5), the FER O Foundation, the Instituto de Salud Carlos Il-Invest iga ci on en Salud(Pl18/01395 and PI21/01019),the Prostate Cancer Foundation(18YOUN19)and the Asociaci on Espanola Contra el Cancer(AECC)(PRY CO 211023 S ERR).J.N.K.is supported by the German Cancer Aid(DECADE,70115166),the German Federal Ministry of Education and Research(PEARL, 01KD2104C; CAMINO, 01EO2101; SWAG, 01KD2215A; TRANSFORM LIVER, 031L0312A; and TANGERINE,O1KT2302 through ERA-NET Trans can), the German Academic Exchange Service (SECAl,57616814),the German Federal Joint Committee(Transplant Kl,01VSF21048),the European Union's Horizon Europeand innovation programme(ODELIA,101057091;and GENIAL,101096312), the European Research Council(ERC;NADIR,101114631) and the National Institute for Health and Care Research(NIHR;NIHR203331)Leeds Biomedical Research Centre.The views expressed are those of the authors and not necessarily those of the NHS, the Nl HR or the Department of Health and Social Care.This work was funded by the European Union. Views and opinions expressed are,however, those of the authors only and do not necessarily reflect those of the European Union. Neither the European Union nor the granting authority canbeheld responsible for them.
R.P.-L. 由 LaCaixa 基金会、CRIS 基金会人才奖 (TALENT19-05)、FER O 基金会、卡洛斯三世健康研究所 (PI18/01395 和 PI21/01019)、前列腺癌基金会 (18YOUN19) 以及西班牙抗癌协会 (AECC) (PRYCO211023SERR) 资助。J.N.K. 获得德国癌症援助 (DECADE, 70115166)、德国联邦教育与研究部 (PEARL, 01KD2104C; CAMINO, 01EO2101; SWAG, 01KD2215A; TRANSFORM LIVER, 031L0312A; 及通过 ERA-NET Transcan 的 TANGERINE, 01KT2302)、德国学术交流中心 (SECAI, 57616814)、德国联邦联合委员会 (TransplantKI, 01VSF21048)、欧盟地平线欧洲创新计划 (ODELIA, 101057091; 及 GENIAL, 101096312)、欧洲研究理事会 (ERC; NADIR, 101114631) 和利兹生物医学研究中心国家健康与护理研究所 (NIHR203331) 的资助。本文观点仅代表作者,不代表英国国家医疗服务体系、NIHR 或卫生与社会保障部立场。本研究由欧盟资助,但所表达观点仅代表作者本人,不一定反映欧盟立场。欧盟及资助机构均不对其承担责任。
Author contributions
作者贡献
All authors contributed substantially to discussion of the content and reviewed and/or edited the manuscript before the submission.R.P.-L., N.G.L.and J.N.K.researched data for the article and wrote the article.
所有作者均对内容讨论做出实质性贡献,并在提交前审阅和/或修改了手稿。R.P.-L. 、N.G.L. 和 J.N.K. 负责文献数据调研与文章撰写。
Competing interests
利益冲突
J.N.K.declares consulting services for Ow kin,Do More Diagnostics,Panakeia,Scailyte, Mind peak and Multiplex Dx;holds shares in Strat if Al GmbH;has received a research grant from GSK;and has received honoraria from Astra Zen eca,Bayer,Eisai,Janssen,MSD,BMS, Roche, Pfizer and Fresenius.R.P.-L.declares research funding by Astra Zen eca and Roche, and participates in the steering committee of a clinical trial sponsored by Roche, not related to this work.All other authors declare no competing interests.
J.N.K. 声明为 Owkin、Do More Diagnostics、Panakeia、Scailyte、Mind Peak 和 Multiplex Dx 提供咨询服务;持有 StratifAI GmbH 股份;获得葛兰素史克 (GSK) 研究资助;并收受阿斯利康 (AstraZeneca)、拜耳 (Bayer)、卫材 (Eisai)、强生 (Janssen)、默沙东 (MSD)、百时美施贵宝 (BMS)、罗氏 (Roche)、辉瑞 (Pfizer) 和费森尤斯 (Fresenius) 的酬金。R.P.-L. 声明获得阿斯利康 (AstraZeneca) 和罗氏 (Roche) 的研究资助,并参与罗氏 (Roche) 赞助的临床试验指导委员会(与本研究无关)。其余作者声明无竞争利益。
Additional information
补充信息
Supplementary information The online version contains supplementary material available at https://doi.org/10.1038/s41568-024-00694-7.
补充信息 在线版本包含补充材料,可在 https://doi.org/10.1038/s41568-024-00694-7 获取。
Peer review information Nature Reviews Cancer thanks the anonymous reviewer(s)fortheir contribution to the peer review of thiswork.
《自然综述·癌症》感谢匿名审稿人对本工作的同行评议所做出的贡献。
Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional a ff li at ions.
出版方注:Springer Nature对出版地图中的管辖权主张和机构从属关系保持中立。
Springer Nature or its licensor (e.g.a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rights holder(s); author selfarchiving of the accepted manuscript version of this articleis solely governed bythe terms of such publishing agreement and applicable law.
Springer Nature 或其许可方 (例如学会或其他合作伙伴) 根据与作者或其他权利持有者签订的出版协议, 对本文享有独家权利; 作者对本文录用稿的自存档仅受此类出版协议条款和适用法律的约束。
