Estimating Uncertainty in Neural Networks for Cardiac MRI Segmentation: A Benchmark Study
Introduction
Cardiac magnetic resonance imaging (MRI) is the gold standard for evaluating cardiac function due to its excellent soft tissue contrast, high spatial and temporal resolution, and non-ionizing radiation . Segmentation of cardiac structures such as the left ventricle cavity, left ventricle myocardium, and right ventricle cavity is required as a first step to quantify clinically relevant imaging biomarkers such as the left ventricle ejection fraction and myocardial mass. Recently, convolutional neural networks (CNNs) have been shown to perform well for automatic cardiac MR image segmentation . However, when using a CNN in an automated image analysis pipeline, it is important to know which segmentation results are problematic and require further manual inspection. This may improve workflow efficiency by focusing on problematic segmentations, avoiding the review of all images and reducing errors in downstream analysis. This problem has been referred to as , and is closely related to the task of anomaly detection or out-of-distribution detection . One approach to solve this problem is to use predictive uncertainty estimates of a segmentation model. The key idea here is that segmentation outputs with low uncertainty are correct while outputs with high uncertainty are problematic. While several studies have attempted to estimate uncertainty in CNNs for medical image segmentation, most of these studies used Monte Carlo (MC) Dropout or Deep Ensembles. However, there are some limitations associated with these methods, which motivates exploration of other algorithms for estimating uncertainty. For example, when using a fixed dropout rate in MC Dropout, the model uncertainty does not decrease as more training data is used. This is potentially problematic since model uncertainty should approach zero in the limit of infinite data . As a result, the dropout rate needs to be tuned depending on the model size and amount of training data . For Deep Ensembles, it is not clear why this method generates well-calibrated uncertainty estimates. In addition, in previous studies, evaluation of these algorithms was mostly limited to correlations between the predictive uncertainty and segmentation accuracy or metrics measuring how uncertainty can be used to improve segmentation . In this work, we compared Bayesian and non-Bayesian approaches and performed a systematic evaluation of epistemic uncertainty for these methods. In particular, we evaluated Bayes by Backprop (BBB), MC Dropout, and Deep Ensembles based on segmentation accuracy, probability calibration, uncertainty on out-of-distribution datasets, and finally, demonstrated the utility of these methods for segmentation quality control.
简介
心脏磁共振成像(MRI)是由于其优异的软组织对比,空间和时间分辨率以及非电离辐射而评估心功能的金标准。作为临床相关的成像生物标志物如左心室喷射分数和心肌肿块,需要诸如左心室腔,左心室心肌和右心室腔的心脏结构的心脏结构,例如左心室腔,左心室心肌和右心室腔。最近,已经显示了卷积神经网络(CNNS)对自动心脏MR图像分割来表现良好。然而,当在自动图像分析管道中使用CNN时,重要的是要知道哪个分段结果是有问题的并且需要进一步的手动检查。这可以通过专注于有问题的分割来提高工作流程效率,避免在下游分析中审查所有图像和减少错误。此问题已被称为,与异常检测或分发检测的任务密切相关。解决这个问题的一种方法是使用分段模型的预测不确定性估计。这里的关键想法是,具有低不确定性的分割输出是正确的,而具有高不确定性的输出是有问题的。虽然几项研究已经尝试估计CNN中的医学图像分割中的不确定性,但这些研究中的大多数研究使用了Monte Carlo(MC)辍学或深度集成。然而,与这些方法有一些局限性,这激发了对其他算法来估算不确定性的探索。例如,在MC丢失中使用固定辍学率时,模型不确定性不会随着使用更多训练数据而减少。这可能是有问题的,因为模型不确定性应该在无限数据的极限下接近零。因此,需要根据培训数据的模型大小和数量来调整辍学率。对于深度合奏,目前尚不清楚为什么此方法产生良好校准的不确定性估计。此外,在先前的研究中,这些算法的评估主要限于预测不确定性和分割精度或度量之间的相关性,测量如何使用不确定性来改善分割。在这项工作中,我们比较了贝叶斯和非贝叶斯方法,对这些方法进行了对认知不确定性的系统评估。特别是,我们通过Backprop(BBB),MC辍学和深度集成的贝叶斯评估了贝叶斯,基于分割精度,概率校准,超出分配数据集的不确定性,最后显示了这些方法进行分割质量控制的效用。
Uncertainty in Neural Networks
Uncertainty in neural networks can be estimated using Bayesian and non-Bayesian methods. Bayesian neural networks (BNNs) provide a theoretical framework for generating well-calibrated uncertainty estimates . In BNNs, we are interested in learning the posterior distribution of the neural network weights instead of a maximum likelihood or maximum-a-posteriori estimate. A challenge in learning BNNs is that integrating over the posterior is intractable in high dimensional space. As such, approximate inference techniques such as stochastic variational inference are commonly used. Examples include variational dropout , MC dropout , Bayes by Backprop , multiplicative normalizing flows , and Flipout . Non-Bayesian methods can also be used to estimate uncertainty in neural networks. Examples include bootstrapping , Deep Ensembles , Temperature Scaling , and Resampling Uncertainty Estimation . Note that in Bayesian methods, uncertainty is learned during training and is tightly coupled to the model structure. In non-Bayesian methods, uncertainty is learned during training or estimated after training. In this work, we focused on modelling epistemic uncertainty using Bayesian and non-Bayesian methods. Epistemic or model-based uncertainty is uncertainty related to model parameters due to the use of a finite amount of training data. For humans, epistemic uncertainty corresponds to uncertainty from inexperienced or non-expert observers. In contrast, aleatoric or data-dependent uncertainty is related to the data itself and corresponds to inter- and intra-observer variability. Epistemic uncertainty is more applicable for segmentation quality control since segmentation outputs with high aleatoric uncertainty may be considered acceptable from a quality control perspective as long as the output matches the expectations from one or more experienced observers.
在神经网络中的不确定性,可以使用贝叶斯和非贝叶斯方法估算神经网络的不确定性。贝叶斯神经网络(BNN)提供了一种用于产生良好校准的不确定性估计的理论框架。在BNN中,我们有兴趣了解神经网络权重的后部分布而不是最大可能性或最大-A-Bouthiori估计。学习BNN的挑战是整合在后部是棘手的高尺寸空间。因此,通常使用诸如随机变分推理的近似推理技术。示例包括变分差,MC丢失,Bayes By BackProp,乘法标准化流程和Flipout。非贝叶斯方法也可用于估计神经网络中的不确定性。示例包括引导,深度集合,温度缩放和重新采样的不确定性估计。请注意,在贝叶斯方法中,在培训期间了解到不确定性,并且紧密地耦合到模型结构。在非贝叶斯方法中,在培训或训练后估计期间了解不确定性。在这项工作中,我们专注于使用贝叶斯和非贝叶斯方法建模认知不确定性。由于使用有限量的培训数据,以外基于史或基于模型的不确定性是与模型参数相关的不确定性。对于人类而言,认知不确定性对应于来自缺乏经验或非专家观察员的不确定性。相比之下,炼体或数据相关的不确定性与数据本身有关并且对应于与观察者间变异性。认知不确定性更适用于分割质量控制,因为由于产量与来自一个或多个经验的观察者的期望相匹配,因此可以认为具有高梯度不确定性的分割输出可能被视为可接受的。
Related Studies
Bayesian neural networks have been used for medical image segmentation tasks; however, most work used MC Dropout to approximate the posterior distribution of the weights and investigations of ways to evaluate the quality of uncertainty have been limited. , , and used a CNN with MC Dropout for brain structure, brain tumour, and brain tumour cavity segmentation. These studies observed positive correlations between segmentation accuracy and uncertainty measures. compared different uncertainty measures in brain lesion segmentation and showed that uncertainty measures can be used to improve lesion detection accuracy. applied MC Dropout on a CNN for cardiac MRI segmentation and showed that training using a Brier loss or cross-entropy loss produced well-calibrated pixel-wise uncertainties, and correcting uncertain pixels can improve segmentation results consistently. used MC Dropout and other non-Bayesian methods to generate skin lesion segmentation and the segmentation uncertainty maps, which were then entered into another neural network to predict the Jaccard index of the segmentation. proposed segmentation quality estimates for aortic MRI segmentation using an ensemble of neural networks and demonstrated improved segmentation accuracy with the use of these estimates. More recently, compared MC Dropout, Deep Ensembles, and auxiliary networks for predicting pixel-wise segmentation errors for two medical image segmentation tasks. In addition to segmentation probability calibration, they examined the overlap between segmentation uncertainty and errors, and the fraction of images which would benefit from uncertainty-guided segmentation correction. In a follow-up work, compared different aggregation methods for uncertainty measures and their performance for segmentation failure detection. Similarly, also compared MC Dropout and Deep Ensembles for CNNs trained with different loss functions in terms of probability calibration and correlation between segmentation accuracy and uncertainty measures. However, these studies did not evaluate other Bayesian methods such as Bayes by Backprop for uncertainty estimation and the performance of these methods on out-of-distribution datasets is unknown. In this work, we evaluated and compared several commonly used uncertainty estimation methods on datasets with various degrees of distortions, which mimic the clinical scenarios. We analyzed the strengths and limitations of these methods on datasets with various distortions, and finally, demonstrated the utility of these algorithms for segmentation quality control.
相关研究
贝叶斯神经网络已用于医学图像分割任务;然而,大多数工作用过的MC辍学以近似重量的后部分布和评估不确定性质量的方式的权重和调查受到限制。 ,并使用具有MC辍学的CNN用于脑结构,脑肿瘤和脑肿瘤腔分割。这些研究观察了分割准确性与不确定性措施之间的正相关。比较了脑病变分割中的不同不确定性措施,并显示了不确定性措施可用于提高病变检测精度。在CNN上的应用MC丢失用于心脏MRI分割,并显示使用Brizer损失或跨熵损失产生良好校准的像素的不确定性,并且校正不确定像素可以一致地改善分割结果。二手MC辍学和其他非贝叶斯方法生成皮肤病变分割和分割不确定性地图,然后进入另一个神经网络以预测分割的Jaccard指数。使用神经网络的集合进行主动脉MRI分割的提出分割质量估计,并通过使用这些估计来证明改善的分割精度。最近,用于比较MC辍学,深度集成和辅助网络,用于预测两个医学图像分割任务的像素 - 明智的分割错误。除了分割概率校准之外,它们还检查了分割不确定度和错误之间的重叠,以及从不确定引导的分割校正中受益的图像的分数。在后续工作中,比较了不同的聚集方法,以了解不确定度量及其分割故障检测的性能。同样,在对不同损失函数训练的CNNS比较了MC辍学和深度集合,在概率校准和分割精度与不确定性措施之间的相关性方面。然而,这些研究没有评估其他贝叶斯方法,如贝叶斯,因为对不确定度估计,并且这些方法对分发外部数据集的性能是未知的。在这项工作中,我们评估并比较了几种常用的不确定性估算方法,在具有各种扭曲的数据集上,模拟了临床情景。我们分析了具有各种扭曲的数据集上这些方法的优势和局限性,最后显示了这些算法进行分割质量控制的效用。
Contributions
While MC Dropout and Deep Ensembles have been used to estimate uncertainty in medical image segmentation, limited studies have investigated other Bayesian methods and a comparison of these methods is currently lacking. To this end, we performed a systematic study of Bayesian and non-Bayesian neural networks for estimating uncertainty in the context of cardiac MRI segmentation. Our contributions are listed as follows: We hope this work will serve as a benchmark for evaluating uncertainty in cardiac MRI segmentation and inspire further work on uncertainty estimation in medical image segmentation.
贡献
虽然MC辍学和深度集成用于估计医学图像分割中的不确定性,但有限的研究研究了其他贝叶斯方法,目前缺乏这些方法的比较。为此,我们对贝叶斯和非贝叶斯神经网络进行了系统研究,以估计心脏MRI分割背景下的不确定性。我们的贡献列出如下:我们希望这项工作将作为评估心脏MRI分割的不确定性的基准,并激发进一步研究医学图像分割中的不确定性估算。
Methods
方法
Bayesian Neural Networks (BNN) 贝叶斯神经网络(BNN)
Given a dataset of $ N $ images $ X= {x_i}$ $ {i=1 \ldots N}$ and the corresponding segmentation $ Y= {y_{i}}_{i=1\ldots N} $ with $ C $ classes, we fit a neural network parameterized by weights $
w $ to perform segmentation. In BNNs, we are interested in learning the posterior distribution of the weights, $ p(w | X, Y)
$ , instead of a maximum likelihood or maximum-a-posteriori estimate. This posterior distribution represents uncertainty in the weights, which could be propagated to calculate uncertainty in the predictions . In addition, BNNs have been shown to be able to improve the generalizability of neural networks . A challenge in learning BNNs is that calculating the posterior is intractable due to its high dimensionality. Variational inference is a scalable technique that aims to learn an approximate posterior distribution of the weights $ q(w) $ by minimizing the Kullback-Leibler (KL) divergence between the approximate and true posterior. This is equivalent to maximizing the evidence lower bound (ELBO) as follows:
给定$ N $的数据集$ X= {x_i}$ $ {i=1 \ldots N}$和相应的分割$ Y= {_{i}}_{i=1\ldots N} $参数化神经网络按权重$
w $执行分段。在BNN中,我们有兴趣学习权重$ p(w | X, Y)
$的后部分布,而不是最大似然或最大-A-Bouthiori估计。该后部分布代表重物中的不确定性,其可以被传播以计算预测中的不确定性。此外,已经证明了BNN能够提高神经网络的普遍性。学习BNNS的挑战是,由于其高维度,计算后部是棘手的。变形推断是一种可伸缩技术,其旨在通过最小化近似和真实后的后退之间的kullback-leibler(kl)发散来学习权重$ q(w) $的近似近似分布。这相当于最大化下限(Elbo)的证据,如下:
$$ q(w) = \argmax_{q(w)}{\mathbb{E}_{q(w)}[\log p(Y| X, w)] - \lambda \cdot \textrm{KL}[q(w) || p(w)]},, $$

where $ \mathbb{E}_ {q(w)}[\cdot] $ denotes expectation over the approximate posterior $ q(w) $ , $ \log p(Y| X,
w) $ is the log-likelihood of the training data with given weights $ w $ , $ p(w) $ represents the prior distribution of $ w $ , and KL[ $ \cdot $ ] is the Kulback-Leibler divergence between two probability distributions weighted by a hyperparameter $ \lambda >0 $ to achieve better performance. State-of-the-art segmentation neural networks such as the U-net formulate image segmentation as pixelwise classification. For each pixel $ x_{i,j} $ in image $ x_i $ , $ i= 1,\ldots N $ , $ j\in\Omega $ , the neural network generates a prediction$ \hat{y}{i,j} $ with probability
$ p(\hat{y}{i,j}=c), c =0, 1, \dots C-1 $ , through softmax activation of the features in the last layer.
其中$\mathbb{E}_ {q(w)}[\cdot] $表示通过近似后后D_d1 q(w) $的预期,$\log p(Y| X,
w) $是给定权重$ w $的训练数据的日志似然性, $ p(w) $表示$ w $的先前分布,KL [$\cdot $]是KLBACK-LEIBLER在由HyperParameter $\lambda >0 $加权的两个概率分布之间的发散,以实现更好的性能。最先进的分割神经网络,例如U-Net将图像分割为PixelW方向分类。对于每个像素$ x_在图像t60_1 $ $ x_i $,$ i= 1,\ldots N $,$ j\in\Omega $,神经网络生成预测$ \hat{y}{i,j} $概率
$ p(\hat{y}{i,j}=c), c =0, 1, \dots C-1 $,通过SOFTMAX激活最后一层中的特征。
Assuming each pixel is independent from other pixels in the image, the log likelihood of the training data in Eq. is given by: where $ y_{i,j} $ is the ground truth label for pixel $ j $ in the $ i^{th} $ image, and [] is the indicator function. In this setting, the log likelihood is the same as the negative cross entropy between the ground truth and predicted segmentation. The prediction $ \hat{y} $ of the BNN on a test image $ x $ is generated by marginalizing out the weights of the neural network, i.e.,
假设每个像素与图像中的其他像素无关,EQ中的训练数据的日志似然性。给出:其中$ y_{i,j} $是$ i^{th} $图像中像素$ j $的地面真理标签,[]是指示灯函数。在此设置中,日志似然与地面真理和预测分割之间的负跨熵相同。通过边缘化神经网络的权重,即
$$ p(\hat{y} | x) = \mathbb{E}_{q(w)}[p(\hat{y}|x, w)], $$
where $ p(\hat{y}|x, w) $ denotes the prediction given a set of weights $ w $ . In the next two sections, we introduce methods for estimating an approximate posterior $ q(w) $ for the weights of a neural network.
其中$ p(\hat{y}|x, w) $表示给定一组权重$ w $的预测。在接下来的两个部分中,我们介绍用于估计近似后的$ q(w) $的方法,用于神经网络的权重。
Bayes by Backprop
One way to parameterize the approximate posterior $ q(w) $ is to use a fully factorized Gaussian. This is sometimes called mean-field variational inference. In a fully factorized Gaussian, each weight $ w $ in $ w $ is independent from other weights and follows a Gaussian distribution with mean $ \mu $ and standard deviation $ \sigma $ . To ensure $ \sigma > 0 $ and training stability, $ \sigma $ is parameterized by a real number $ \rho $ , i.e., $ \sigma =
\textrm{softplus}(\rho) = \ln(1+e^\rho) $ . The prior distribution $ p(w) $ is usually chosen as a fully factorized Gaussian with mean $ \mu_{prior}I $ and standard deviation $ \sigma_{prior}I $ , i.e., $ p(w) = \mathcal{N}(\mu_{prior}I, \sigma_{prior}\mathcal{N}0 $ , where $ I $ represents the identity matrix. Gradient updates can be performed using the reparameterization trick. The training procedure is known as Bayes by Backprop (BBB) and described below: After training, each weight can be sampled from $ \mathcal{N}(\mu, \sigma) $ , which is then used to generate the predicted segmentation using Eq. .
Bayes by BackProp
将近似后$ q(w) $的一种方法是使用完全分解的高斯。这有时被称为卑鄙场变分推理。在一个完全分解的高斯,$ w $中的每个权重$ w $独立于其他权重,遵循具有平均$\mu$和标准偏差$\sigma $的高斯分布。为了确保$\sigma > 0 $和训练稳定性,$\sigma $由实数$\rho $,即$\sigma =
\textrm{softplus}(\ln(1+e^\rho) $。先验分布$ p(w) $通常被选择作为一个完全因式分解高斯均值$\mu_{prior}I $和标准偏差$\sigma_{prior}I $,即$ p(w) = \mathcal{N}(\mu_{prior}I, \sigma_{prior}\mathcal{N}0 $,其中$ I $表示身份矩阵。可以使用Reparameterization技巧执行渐变更新。培训程序被回溯(BBB)称为贝叶斯(BBB)并如下所述:在训练之后,可以从$\mathcal{N}(\mu, \sigma) $中采样每个重量,然后使用EQ生成预测的分段。 。
MC Dropout
MC Dropout (MCD) is another commonly used method for learning BNNs because it is straightforward to implement and does not require additional parameters or weights. MCD can be interpreted as choosing the approximate posterior distribution $ q(w) $ to be a mixture of two Gaussians with minimal variances, e.g., one at $ 0 $ and the other at the weight $ w $ . Dropout is applied during training and testing to sample weights from $
q(w) $ . In this method, the dropout rate is a hyperparameter chosen empirically based on a validation dataset. The dropout rate defines the amount of uncertainty in the weights and is fixed throughout network training and testing.
MC丢失
MC丢失(MCD)是另一个用于学习BNN的常用方法,因为它很简单地实现并且不需要额外的参数或权重。 MCD可以被解释为选择近似后部分布$ q(w) $,其具有最小差异的两个高斯的混合,例如,在重量$ w $处为$ 0 $和另一个。在训练和测试期间应用辍学,以从$
q(w) $进行采样权重。在此方法中,辍学率是基于验证数据集的凭证选择的超级计。辍学率定义了重量中的不确定性量,并在整个网络培训和测试中得到了固定。
Deep Ensembles
In addition to BNNs, we characterized and evaluated uncertainty estimates using an ensemble of neural networks. This is also known as Deep Ensembles . Deep Ensembles consist of multiple neural networks trained using the same data (or different subsets of the same data) with different random initializations. Combination of the models in an ensemble has been shown to produce well-calibrated probabilities in computer vision tasks, and the variability in the models can be used to calculate predictive uncertainty. This non-Bayesian approach was inspired by the idea of bootstrapping, where stochasticity in the sampling of the training data and training algorithm define model uncertainty. This approach differs from Bayesian methods since it does not require approximation of the posterior distribution of the weights.
除了BNN之外,我们还使用神经网络的集合来表征和评估不确定性估计,### Deep Ensembles
。这也被称为深度集成。深度集合由使用具有不同随机初始化的相同数据(或不同数据的不同子集)训练的多个神经网络。已经显示了集合中模型的组合在计算机视觉任务中产生了良好的校准概率,并且可以使用模型的可变性来计算预测性不确定性。这种非贝叶斯方法受到自动启动的启发,其中培训数据和培训算法采样的随机性定义了模型不确定性。这种方法与贝叶斯方法不同,因为它不需要近似重量的后部分布。
Algorithm Evaluation
We evaluated the uncertainty estimation algorithms based on three aspects: (1) segmentation accuracy and probability calibration, (2) uncertainty on out-of-distribution images, and (3) application of uncertainty estimates for segmentation quality control. The purposes of these evaluations are as follows: (1) We show that Bayesian neural networks can provide segmentation accuracies that are similar or higher than plain or point estimate neural networks. In addition, predicted segmentation probabilities should be well-calibrated, i.e., a pixel with a predicted probability of 60% belonging to the myocardium is truly 60%. From a frequentist perspective, this means that out of all predictions with 60% probability, 60% of the predictions are correct. (2) We measure segmentation uncertainty on out-of-distribution images to validate that the uncertainty measures perform as expected, i.e., uncertainty should increase when test images are substantially different from training datasets. (3) We expect uncertainty measures to be useful in identifying problematic segmentations that require manual editing. We first discuss metrics for segmentation accuracy and probability calibration and then present methods to quantify predictive uncertainty using uncertainty measures.
算法评估
我们评估了基于三个方面的不确定性估计算法:(1)分割精度和概率校准,(2)在分配图像外的不确定性,以及(3)对分割质量控制的不确定性估计的应用。这些评估的目的如下:(1)我们表明贝叶斯神经网络可以提供类似或高于普通或点估计神经网络的分段精度。另外,预测的分割概率应该是良好的校准,即,具有预测概率的像素,其属于心肌的60%的像素是真正的60%。从常见的角度来看,这意味着超过了60%概率的预测,60%的预测是正确的。 (2)我们测量分发图像的分割不确定度,以验证不确定性措施,即当测试图像与训练数据集的基本不同时,不确定性应该增加。 (3)我们预计不确定性措施可用于识别需要手动编辑的问题分割。我们首先讨论分割精度和概率校准的指标,然后使用不确定性措施来定量预测性不确定性的方法。
Segmentation Accuracy
Let $ R_a $ and $ R_m $ be the automated and manual segmentation regions, respectively. We calculated the algorithm segmentation accuracy using Dice similarity coefficient, average symmetric surface distance (ASSD), and Hausdorff distance (HD). measures the overlap between two segmentations and is given by: quantifies the average distance between the contours of two segmentation regions and is given by: where $ \abs{\partial R_{a}} $ represents the number of points on contour $ \partial R_{a} $ and $ d(p,\partial R_m) $ represents the shortest Euclidean distance from point $ p $ to contour $ \partial R_m $ ; $ \abs{\partial R_m} $ and $ d(p,\partial R_a) $ are defined in the same manner. is the maximum distance between two contours and is calculated as:
分段精度
让$ R_a $和$ R_m $分别为自动和手动分段区域。我们使用骰子相似系数,平均对称表面距离(ASSD)和Hausdorff距离(HD)计算算法分割精度。测量两个分段之间的重叠,并给出:定量两个分割区域的轮廓之间的平均距离,并给出:其中$\abs{\partial R_{a}} $表示轮廓上的点数$\partial R_{a} $和$ d(p,\partial R_m) $表示的点数从点$ p $到轮廓$\partial R_m $的最短欧几里德距离; $\abs{\partial R_m} $和$ d(p,\partial R_a) $以相同的方式定义。是两个轮廓之间的最大距离,并计算为:
Probability Calibration
These metrics measure how closely the neural network segmentation probabilities match the ground truth probabilities generated from manual segmentation on a per-pixel basis. Following the notation in Sec. bnn-intro, we let $ \hat{y}_j $ and $ y_j $ denote the prediction and ground truth label of pixel $ j $ in a given image, $ j\in\Omega $ , respectively. measures how well the learned model fits the observed (testing) data. Note that NLL is sensitive to tail probabilities; that is, a model that generates low probability for the correct class is heavily penalized. NLL is calculated as: measures the mean squared error between the predicted and ground truth probabilities and is given by:
概率校准
这些度量标准测量神经网络分割概率的敏捷程度与每个像素的手动分段产生的地面真相概率匹配。在秒中的符号之后。 BNN-intro,我们让$\hat{y}_j $和$ y_j $分别表示给定图像中的像素$ j $的预测和地形标签,$ j\in\Omega $。测量学习模型适合观察到的(测试)数据的程度。请注意,NLL对尾部概率敏感;也就是说,为正确阶级产生低概率的模型严重惩罚。 nll计算为:测量预测和地面真实概率之间的平均平均误差,并提供:
Predictive Uncertainty Measures 预测性不确定性测量
Predictive uncertainty measures can be calculated from the neural network predictions to indicate the degree of certainty of the output. These can be calculated per pixel or per structure/class. Pixelwise uncertainty measures are motivated by information theory. These values are calculated per pixel and averaged across all pixels in the image if an image-level measure is required. In this work, we used the following pixelwise uncertainty measures: - {Predictive Entropy} measures the spread of probabilities across all the classes in the mean prediction and is given by:
预测性不确定性措施可以从神经网络预测计算,以指示输出的确定性程度。这些可以每像素或每个结构/类计算。 PixelWive不确定性措施是通过信息理论的动机。如果需要图像级度量,则每个像素计算每个像素并在图像中的所有像素中的平均值。在这项工作中,我们使用以下PixelWise不确定性措施: - {预测熵}在平均预测中的所有类中测量概率的传播,并通过
$$ \sum_{j \in \Omega} \sum_{c=0}^{C-1} \left[-p \left(\hat{y}_j = c \right)\log p \left( \hat{y}_j = c \right)\right] $$
- {Mutual Information (MI)} measures how different each sample is from the mean prediction and is calculated as:
- MI 测量每个样本的不同从平均预测中的不同,并且计算为:
- $$ \mathbb{E}{q(\mbf{w})} & \left[ \sum{j\in \Omega} \sum_{c=0}^{C-1} , p \left ( \hat{y}j = c | \mbf{w} \right ) , \log p \left ( \hat{y}j = c|\mbf{w} \right)- \right . & \left . \sum{j\in \Omega} \sum{c=0}^{C-1} , p \left(\hat{y}_j = c \right), \log p \left(\hat{y}_j = c \right), \right ] \ $$
$
where p \left(\hat{y}_j = c|\mbf{w} \right) denotes the prediction given a set of weights w . MI is high if there are samples with both high and low confidence, and is low if all samples have low confidence or high confidence. Structural uncertainty measures were proposed specifically for image segmentation . Here, we define two structural uncertainty measures, which quantify how different each structure is among the prediction samples in terms of Dice and ASSD. We expect structural uncertainty measures to better align with common segmentation accuracy metrics because of their focus on global image-level uncertainty. - {Dice\textsubscript{WithinSamples}} =\frac{1}{T} \sum_{i=i}^{T} \textrm{Dice}(\bar{S}, S_{i}) - {ASSD\textsubscript{WithinSamples}} =\frac{1}{T} \sum_{i=i}^{T} \textrm{ASSD}(\bar{S}, S_{i}) where $ \bar{S} $ is the mean predicted segmentation and $ S_{i}, i \in{1\ldots T} $ , are individual predication samples from the neural network.
其中 p \ left(\ hat {y} _j = c | \ mbf {w} \右)表示给出的预测一套重量 w 。如果有高度和低置信度的样品,MI很高,如果所有样品都具有低置信度或高信心,则低。专门针对图像分割提出了结构性不确定性措施。在这里,我们定义了两个结构不确定性措施,该措施量化了各种结构在骰子和截例方面的预测样本中的不同程度。我们预计结构不确定度量措施与常见的分割准确性指标更好地对齐,因为它们专注于全球图像级不确定性。 - {dice \ textsubscript {withinsamples}} = \ frac {1} {t} \ sum_ {i = i} ^ {t} \ textrm {dice}(\ bar {s},s_ {i}) - { assd \ textsubscript {withinsamples}} = \ frac {1} {t} \ sum_ {i = i} ^ {t} \ textrm {assd}(\ bar {s},s_ {i}) where $\bar{S} $是平均预测分割和$ S_{i}, i \in{1\ldots T}$,是来自神经网络的单独预测样本。
Datasets 数据集
UK BioBank (UKBB)
The UKBB dataset consists of images from 4845 healthy volunteers. For each subject, 2D cine cardiac MR images were acquired on a 1.5T Siemens scanner using a bSSFP sequence under breath-hold conditions with ECG-gating (pixel size = 1.8-2.3 mm, slice thickness = 8 mm, number of slices = $ \sim $ 10, number of phases = $ \sim $ 50). Manual segmentation of the left ventricle blood pool (LV), left ventricle myocardium (Myo), and right ventricle (RV) was performed on the end-diastolic (ED) and end-systolic (ES) phases by one of eight observers followed by random checks by an expert to ensure segmentation quality and consistency. The dataset was randomly split into 4173, 103, and 569 subjects for training, validation, and testing, respectively. These numbers were chosen for convenience.
英国biobank(UKBB)
UKBB数据集由4845个健康志愿者的图像组成。对于每个受试者,在1.5T Siemens扫描仪上使用BSSFP序列在呼吸保持条件下使用ECG-Gating(像素尺寸= 1.8-2.3mm,切片厚度= 8mm,切片数量= $)在1.5T西门子扫描仪上获得2D Cine心脏MR图像。\sim $ 10,阶段数= $\sim $ 50)。左心室血液池(LV),左心室心肌(MyO)和右心室(RV)的手动分割是在八个观察者中的一个接下来的末端舒张末期(ED)和末端收缩期(ES)阶段进行的由专家随机检查,以确保分割质量和一致性。数据集分别随机分为4173,103和569个受试者,分别进行培训,验证和测试。为方便起见,选择这些数字。
Automated Cardiac Diagnosis Challenge (ACDC)
The ACDC dataset consists of 100 patients with one of five conditions: normal, myocardial infarction, dilated cardiomyopathy, hypertrophic cardiomyopathy, and abnormal right ventricle. 2D Cine MR images were acquired using a bSSFP sequence on a 1.5T/3T Siemens scanner (pixel size = 0.7-1.9 mm, slice thickness = 5-10 mm, number of slices = 6-18, number of phases = 28-40). Manual segmentation was performed at the ED and ES phases with approval by two experts. This dataset was used for testing only.
自动心脏诊断挑战(ACDC)
ACDC数据集由100名患者组成,其中五种病症中的一个:正常,心肌梗死,扩张心肌病,肥厚性心肌病,和异常右心室。使用BSSFP序列在1.5T / 3T Siemens扫描仪上获得2D Cine MR图像(像素尺寸= 0.7-1.9 mm,切片厚度= 5-10mm,切片数= 6-18,相位数= 28-40 )。手动分割在ED和ES阶段进行,并通过两位专家批准进行。此数据集仅用于测试。
Training Details
We used a plain 2D U-net for MC Dropout, BBB, and Deep Ensembles. The plain U-net consisted of 10 layers with $ 3\times3 $ filters and 2 layers with $ 1\times1 $ convolutions followed by a softmax layer. The number of filters ranged from 32 to 512 from the top to the bottom layers. For MCD, we added dropout on all layers or only on the middle layers of the U-net with different dropout rates: 0.5, 0.3, 0.1. These settings effectively tuned the amount of uncertainty in the model. For BBB, we experimented with different standard deviations of the prior distributions: $ \sigma_{prior}= $ 0.1 or 1.0 and different weights for the KL term: $ \lambda= $ 0.1, 1.0, 10, 30. These are commonly used hyperparameters in the literature . For both methods, the final prediction was obtained by averaging the softmax probabilities of 50 samples. For Deep Ensembles, we trained 10 plain U-net models separately using all of the training data with different random initializations and averaged the softmax probabilities of the 10 models. For each method, we saved models with the lowest NLL on the validation dataset since NLL is directly related to segmentation accuracy and probability calibration. For preprocessing, the input images were cropped by taking a region of 160x160 pixels from the center of the original images. Then, image intensities were normalized by subtracting the mean and dividing by the variance of the entire training dataset. Experiments were repeated 5 times (except for Deep Ensembles) and the results were averaged. Data augmentation was performed, including random rotation (-60 to 60 degrees), translation (-60 to 60 pixels), and scaling (0.7 to 1.3 times). These models were trained using the Adam optimizer for 50 epochs. The initial learning rate was set to 1e-4 which was decayed to 1e-5 after 30 epochs.
训练详细信息
我们使用了一个普通的2D U-Net for MC辍学,BBB和Deep Ensembles。普通U-NET由10层,具有$ 3\times3 $滤波器和2层,具有$ 1\times1 $卷积,然后是Softmax层。从顶部到底层的滤光片的数量范围为32到512。对于MCD,我们在所有层上添加了丢失或仅在U-Net的中间层上,不同的辍学率:0.5,0.3,0.1。这些设置有效地调整了模型中的不确定性量。对于BBB,我们尝试了现有分布的不同标准偏差:$\sigma_{prior}= $ 0.1或1.0以及KL项的不同权重:$\lambda= $ 0.1,1.0,10,30。这些是常用的超参数文学 。对于这两种方法,通过平均50个样品的软MAX概率获得最终预测。对于深度合奏,我们使用不同随机初始化的所有培训数据分别培训了10个普通U-Net模型,并平均10个型号的SoftMax概率。对于每种方法,由于NLL与分段精度和概率校准直接相关,因此我们将使用最低NLL的MLL上保存模型。为了预处理,通过从原始图像的中心拍摄160x160像素的区域来裁剪输入图像。然后,通过减去整个训练数据集的均值和除以整个训练数据集的方差来归一化图像强度。重复实验5次(深度合奏除外),结果平均。执行数据增强,包括随机旋转(-60至60度),转换(-60到60像素)和缩放(0.7到1.3次)。这些模型使用ADAM Optimizer培训50个时期。初始学习速率设定为1E-4,在30个时期之后衰减至1E-5。
Experiments and Results 实验和结果
For BBB, $ \lambda=10 $ and $ \sigma_{prior}=0.1 $ achieved the best validation negative log likelihood. For MC Dropout, adding dropout on the middle layers with a dropout rate of 0.1 (MCD-0.1) performed the best. We also reported results for MC Dropout with a dropout rate of 0.5 on the middle layers (MCD-0.5) for comparison since this setting is commonly used in the literature.
对于bbb,$\lambda=10 $和$\sigma\sigma\sigma\sigma\sigma{prior}=0.1 $实现了最佳验证负对数似然。对于MC辍学,辍学率的中间层上的辍学率最佳地表现为0.1(MCD-0.1)。我们还报告了MC辍学的结果,中间层上的辍学率(MCD-0.5),因为该设置常用于文献中。
Segmentation Accuracy and Probability Calibration
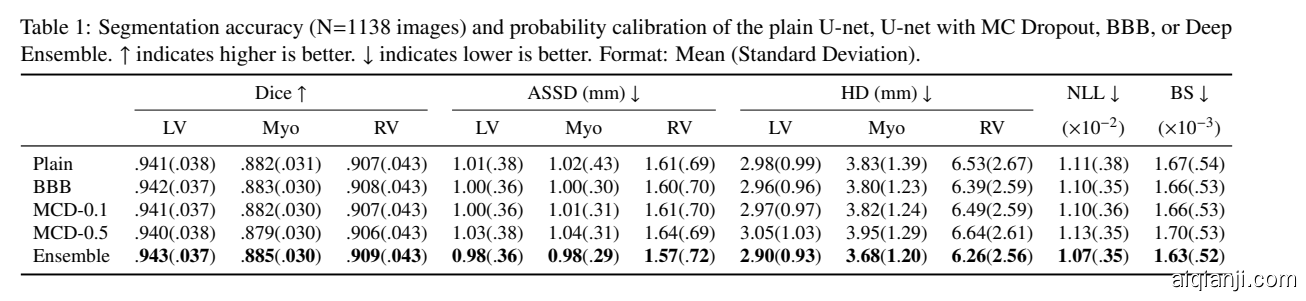
The segmentation accuracy and probability calibration of the U-net trained with various methods are shown in Table segmentation_performance. Deep Ensembles performed the best in terms of segmentation accuracy and probability calibration. This was followed by BBB and MCD-0.1, which were comparable to the plain U-net. MCD-0.5 performed slightly worse than other models in terms of segmentation accuracy and probability calibration. These results indicate that Bayesian approaches or Deep Ensembles can yield similar, if not better, segmentation results compared to a plain U-net while providing uncertainty estimates at the same time.
分段精度和概率校准
具有各种方法的U-Net培训的分割精度和概率校准在表段中显示为表Semonation_Performance。在分割精度和概率校准方面,深度集合是最佳的。这是BBB和MCD-0.1,其与普通U-Net相当。根据分割精度和概率校准,MCD-0.5在其他模型中略差差。这些结果表明,贝叶斯方法或深度集合可以屈服于相似,如果不是更好的,分段结果与普通U-Net相比同时提供不确定性估计。
Uncertainty on Distorted Images
In order to validate uncertainty measures as indicators of out-of-distribution datasets, we applied the trained models to carefully generated images with various magnitudes of distortions, including: - adding Rician noise, as found in MR images , with magnitudes ranging from 0.05 to 0.10 (images intensities ranged from 0 to 1), - Gaussian blurring with a standard deviation of 1.0 to 4.0 pixels, and - deforming or stretching around the cardiac structures. Note that these distortions were not applied as part of data augmentation during training and these images were not seen by the neural network. Figure distortions_example shows examples of the distorted images. We expected decreased segmentation accuracy and increased predictive uncertainty in images with greater magnitudes of distortions.
对扭曲图像的不确定性
,以验证不确定性措施作为外部分发数据集的指标,我们应用训练型模型以仔细生成具有各种扭曲的图像,包括: - 添加瑞典噪声,如MR图像中所说,幅度范围为0.05至0.10(图像强度范围为0至1), - 高斯模糊,标准偏差为1.0至4.0像素,围绕心脏结构变形或拉伸。请注意,这些失真未作为训练期间数据增强的一部分应用,并且神经网络未看到这些图像。 Figure Distortions_example显示扭曲图像的示例。我们预期的分割准确性降低,并提高了图像的预测性不确定性,具有更大的扭曲量大。
Trends with Increasing Distortions
Figureall_distortion_boxplots and Supplementary Table 1 show that segmentation accuracy decreased as the magnitude of the distortions (noise, gaussian blur, stretch) was increased. This is expected since these types of distortions were not seen during training and increasing the magnitude of the distortions resulted in greater differences with the original training datasets. In addition, we observed that the predictive uncertainty increased (i.e., higher predictive entropy, mutual information, ASSD, and lower Dice) with increasing magnitude of distortions but decreased after a certain threshold, as shown in Figureall_distortion_boxplots. This was the case for Deep Ensembles, BBB, and MC Dropout on images with blurring and noise distortions but not with stretching. For example, for BBB, the median predictive entropy for images with slight, moderate, and large additional noise was $ 1.66 \times 10^{-2} $ , $ 1.95 \times 10^{-2} $ , and $ 1.23 \times
10^{-2} $ , respectively. Similarly, the median ASSD for the BBB model was 0.40, 3.40, and 0.71 mm for images with slight, moderate, and large amount of blurring, respectively (See Supplementary Table 3 for more details). Figure noise_distortion_image shows examples of segmentation results and the corresponding predictive entropy and mutual information. While the increasing predictive uncertainty associated with increasing magnitude of distortions was expected, the decrease in predictive uncertainty after a threshold in cases of noise and blurring distortions was not expected. Specifically, for images that were highly distorted, all pixels were being classified as background with low uncertainty (Figureall_distortion_boxplots, bottom row). Although this seems correct when only given the labeling choice of background, LV, Myo, and RV, we argue that the distorted pixels are markedly different from the background pixels in the training images and therefore, should have high uncertainty nonetheless. This is a limitation of all the uncertainty models tested and may be improved using more expressive posteriors. Another observation is that the uncertainty measures began to fail/decrease when dramatic segmentation errors occured (Figureall_distortion_boxplots). This suggests that other heuristics or algorithms such as those presented in can be used to complement the uncertainty measures when trying to detect inaccurate segmentation. For example, segmentation with a non-circular LV blood pool or a blood volume $ < $ 50 mL is highly problematic and may indicate poor segmentation.


增加扭曲幅度后的观测
FIGURALL_DISTOL_BOXPLOT和补充表1表明,随着扭曲的幅度(噪音,高斯模糊,拉伸)增加,分割精度降低。这是预期的,因为在训练期间没有看到这些类型的扭曲,并且增加扭曲的幅度导致与原始训练数据集更大的差异。此外,我们观察到,预测性不确定性增加(即,更高的预测熵,互信息,截图和下骰子),随着较大的差异,但在某个阈值之后降低,如图所示。对于具有模糊和噪声扭曲的图像上的深度集成,BBB和MC辍学,这是如此。例如,对于BBB,具有轻微,中等和大额外噪声的图像的中值预测熵是$ 1.66 \timesT265_2{-2} $,$ 1.95 \times 1.95 {-2} $,以及$ 1.23 \times
10^\times
10^{-2} $。同样,BBB型号的中位数分别为0.40,3.40和0.71 mm,用于微小,中等和大量模糊的图像(有关更多细节,请参见补充表3)。图诊断_distorgatorrage_image显示分割结果的示例和相应的预测熵和相互信息。虽然预期噪声和模糊畸变的阈值之后的预测性不确定性增加了与增加的扭曲程度增加的预测性不确定性。具体地,对于高度扭曲的图像,所有像素都被归类为具有低不确定性的背景(FigureAll_Distorgatorration_Boxplots,底行)。虽然这似乎是正确的,但当鉴于背景,LV,MyO和RV的标签选择时,我们认为失真的像素与训练图像中的背景像素显着不同,因此,应该具有高不确定性。这是对所有测试的不确定性模型的限制,并且可以使用更多富有表现力的前后改善。另一个观察是,在发生戏剧性分割错误时,不确定性措施开始失败/减少(Figureall_distorgatorration_boxplots)。这表明,其他启发式或算法,如诸如所呈现的那些算法可以用于在试图检测到不准确的细分时补充不确定性措施。例如,具有非圆形LV血液池或血液体积$ < $ 50mL的分割非常有问题,并且可以表示分割不良。
Comparison between Deep Ensemble, BBB and MC Dropout
In terms of segmentation accuracy and probability calibration, BBB was more robust to noise distortions compared to the other methods. Specifically, BBB showed higher LV Dice, higher LV ASSD, lower NLL, and lower BS on images with greater noise distortions (Figureall_distortion_boxplots, noise = 0.07, 0.08, 0.10 and Supplementary Tables 1 and 2, degree of distortion = 2, 3, 4). Deep Ensembles was more robust to the blurring and stretching distortions than the other methods. Deep Ensembles showed higher segmentation accuracy in cases with greater blurring and stretching distortions compared to the other methods (Figureall_distortion_boxplots and Supplementary Table 1, degree of distortion = 3, 4). In terms of predictive uncertainty measures, BBB was more robust to the noise distortion while both Deep Ensembles and BBB were more resistant to the blurring compared to the other methods. For example, the mutual information, Dice, and ASSD uncertainty measures peaked at a noise distortion with degree=3 for BBB. In contrast, the uncertainty measures peaked at a noise distortion with degree=2 for Deep Ensemble, MCD-0.1, and MCD-0.5. Comparing the uncertainty on images with heavy noise distortions (degree of distortion = 3, 4), BBB showed the highest predictive uncertainty. This was followed by MCD-0.5 while Deep Ensembles and MCD-0.1 showed the lowest predictive uncertainties. For images that were heavily blurred, Deep Ensembles and BBB showed the highest predictive uncertainty. In cases of images with stretching distortions, all methods showed fairly similar predictive uncertainty across all degrees of stretching. Having high predictive uncertainties on heavily distorted images is desirable since they have lower segmentation accuracy. Overall, BBB showed good predictive uncertainties on all of the tested distortions while Deep Ensemble showed good predictive uncertainties only in cases of blurring and stretching distortions. MCD-0.1 and MCD-0.5 showed results comparable to BBB and Deep Ensembles only on the stretched datasets.
Deep Ensemble, BBB and MC Dropout的比较
之间的比较,与其他方法相比,BBB对噪声扭曲更加坚固。具体地,BBB在具有更大噪声失真的图像上显示出更高的LV骰子,更高的LV ASSD,更低的NLL和下BS(Figureall_Distorgatory_BoxPlots,噪声= 0.07,0.08,0.07,0.08,0.10和补充表1和2,失真程度= 2,3,4 )。深度集合对模糊和拉伸扭曲比其他方法更加强大。与其他方法相比,在更大的模糊和拉伸扭曲的情况下,深度集成精度在更大的模糊和拉伸扭曲(Figurealll_distorgatory_boxpots和补充表1的情况下,失真= 3,4)。就预测性不确定性措施而言,与其他方法相比,BBB对噪声失真更加强劲地对噪声失真更耐受模糊更耐受模糊。例如,相互信息,骰子和ASSD不确定性测量以BBB的度= 3的噪声失真达到峰值。相反,不确定度测量以深度= 2的噪声失真达到峰值,用于深集合,MCD-0.1和MCD-0.5。比较具有重度噪声扭曲的图像的不确定性(畸变程度= 3,4),BBB显示出最高的预测性不确定性。接下来是MCD-0.5,而深度集成和MCD-0.1显示出最低的预测性不确定性。对于严重模糊的图像,深度合奏和BBB显示出最高的预测性不确定性。在具有拉伸扭曲的图像的情况下,所有方法在所有伸展程度上都显示出相同的预测性不确定性。由于它们具有较低的分割精度,因此希望具有高预测性的不确定性。总体而言,BBB对所有测试扭曲显示出良好的预测性不确定因素,而深度集合仅在模糊和拉伸扭曲的情况下显示出良好的预测性不确定性。 MCD-0.1和MCD-0.5显示出与BBB和深度仅在拉伸的数据集上相当的结果。
Uncertainty on Dataset Shift
To further validate the uncertainty measures, we applied the models trained on the UKBB dataset to a different dataset - the ACDC dataset. The ACDC dataset consists of images with pathologies and image contrast distinctly different from the UKBB training dataset. Because of the dataset shift, we expect decreased segmentation accuracy and increased predictive uncertainty on the ACDC dataset compared to the UKBB test dataset. Figure metrics_correlations shows the segmentation accuracy and uncertainty metrics of the U-net models tested on the ACDC dataset. As expected, we observed decreased segmentation accuracy and increased predictive uncertainty compared to the UKBB test dataset. In terms of segmentation accuracy and probability calibration, the methods from the best to the worst are: Deep Ensemble, BBB, MCD-0.1, Plain U-net, and MCD-0.5, as shown in Figure metrics_correlations and Supplementary Table 4. The methods with the highest to the lowest uncertainties are: MCD-0.5, Deep Ensemble, BBB, and MCD-0.1, as shown in Figure metrics_correlations and Supplementary Table 5. Having high uncertainty on a shifted dataset is desirable, especially when the segmentation performance is poor. Overall, both Deep Ensembles and BBB showed good segmentation accuracy and predictive uncertainty in cases of shifted dataset.
在数据集shift上的不确定性
进一步验证了不确定性措施,我们将培训的模型应用于UKBB数据集的模型到另一个数据集 - ACDC数据集。 ACDC数据集由具有病例和图像的图像组成,对比与UKBB训练数据集明显不同。由于数据集移位,我们预计与UKBB测试数据集相比,我们预计将降低分割精度并增加了ACDC数据集上的预测性不确定性。图METRICS_CORRELATIONS显示了在ACDC数据集上测试的U-NET模型的分段精度和不确定性指标。正如预期的那样,与UKBB测试数据集相比,我们观察到的分割准确性和预测性不确定性增加。在分割精度和概率校准方面,来自最坏的方法是:深度集成,BBB,MCD-0.1,普通U-Net和MCD-0.5,如图Metrics_correlations和补充表4所示。该方法最高到最低的不确定性是:MCD-0.5,Deep Ensemble,BBB和MCD-0.1,如图Metrics_correlations和补充表5所示。需要在换档数据集上具有高不确定性,特别是当分段性能较差时。总体而言,深度集成和BBB在移位数据集的情况下,BBB都显示出良好的分割准确性和预测性不确定性。
Correlations between Uncertainty and Segmentation Accuracy
To demonstrate the potential utility of uncertainty measures, we evaluated the Spearman rank correlation between uncertainty measures and segmentation accuracy. We used the rank correlation instead of linear correlation to reduce effects of a potential non-linear relationship between the two quantities. Figure metrics_correlations shows the Spearman rank correlation between the uncertainty measures and the ASSD on the UKBB and ACDC test datasets. In the case of training and testing on the UKBB dataset (UKBB $ \rightarrow $ UKBB), the uncertainty measure that correlated the most with ASSD was ASSD(Spearman correlation $ =\left[0.58,
0.69\right] $ ). This is not surprising since ASSD calculation was similar to ASSD. Similar observations were obtained for training on the UKBB dataset and testing on the ACDC dataset (UKBB $ \rightarrow $ ACDC) although the correlations were slightly lower (Spearman correlation $ =\left[0.44, 0.66\right] $ ). These correlations suggest that the ASSD uncertainty measure is useful in predicting the segmentation quality when ground truth is not available. Figureuncertainty_measures shows representative segmentation results, posterior prediction samples, and the structural uncertainty measures for RV. The images with poor segmentation (toward the right side) had greater uncertainty as measured by Dice and ASSD. Note that posterior prediction samples are not meant to reflect human inter- or intra-observer variability but rather show what the network has learned from the given data.


不确定和分割精度之间的相关性
展示不确定性措施的潜在效用,我们评估了不确定度量和分割精度之间的矛盾等级相关性。我们使用秩相关而不是线性相关性,以减少两种数量之间的潜在非线性关系的影响。图METRICS_CORRELATIONS显示了SPEARMAN位于UKBB和ACDC测试数据集上的不确定性度量和ASSD之间的秩相关性。在UKBB数据集上培训和测试的情况下(UKBB $\rightarrow $ UKBB),与ASSD相关的不确定性措施是ASSD(Spearman相关$ =\left[0.58,
0.69\right] $)。由于ASSD计算类似于ASSD,这并不奇怪。尽管相关性略低(Spearman相关性$ =\left[0 =\left[0.44, 0.66\left[0.44, 0.66\right] $),但获得了在UKBB数据集上进行培训和在ACDC数据集中进行培训并在ACDC数据集上进行测试(UKBB $\rightarrow $ ACDC)。这些相关性表明,ASSD不确定度量可用于预测地面真理不可用的分割质量。图城图绘图显示了代表性分割结果,后预测样本以及RV的结构不确定性措施。分割不良(朝向右侧)的图像具有更大的不确定性,通过骰子和截例测量。请注意,后部预测样本并不意味着反映人类或观察者内的帧内变异性,而是显示网络从给定数据中了解的内容。
Uncertainty for Segmentation Quality Control
In this section, we explored the use of predictive uncertainty estimates to flag potentially problematic segmentations that require manual review. We view this task as a classification problem and we aim to use uncertainty measures to classify segmentations as either good or poor. Having experts to manually determine whether an automated segmentation is good or not for a large dataset is time consuming and adds observer noise. Instead, we used thresholds on segmentation accuracies to determine good or poor automated segmentation. Based on our experience and discussions with our clinical collaborators, we think that contour or surface distance is more indicative of inaccurate segmentations. As such, for each method, the predicted segmentation was considered as poor when the ASSD between the prediction and ground truth is greater than the ASSD of inter-observer manual segmentations. We used inter-observer ASSD of 1.17 mm for LV, 1.19 mm for Myo, and 1.88 mm for RV, based on a recent relatively large-scale study . We then evaluated how well the ASSD uncertainty measure could identify poor segmentations. To utilize the ASSD uncertainty measure, a threshold can be set such that any segmentation with uncertainty above the threshold is flagged for manual review. This would hopefully result in a decreased number of poor segmentation in the dataset. Figureprecision_plots_assd shows the fraction of images with poor segmentation remaining in the dataset and the fraction of images flagged for manual correction when the uncertainty thresholds were varied, i.e., (positives - true positives) vs(true positives + false positives), where positive represents poor segmentation.
在本节中分割质量控制
的不确定性,我们探讨了使用预测的不确定性估计,以销售需要手动审查的潜在问题的细分。我们将此任务视为分类问题,我们的目标是使用不确定性措施将分段分类为好或差。有专家可以手动确定自动分割是否好或不为大型数据集是耗时的,并增加观察者噪声。相反,我们在分割精度上使用了阈值来确定自动分割的良好或差。根据我们的经验和讨论我们的临床合作者,我们认为轮廓或表面距离更为指示不准确的细分。因此,对于每种方法,当预测和地面事实之间的ASSD大于观察者间手动分割的ASSD时,预测的分割被认为是差的。基于最近的近期大规模研究,我们使用17 mm的Observer ASSD为1.17 mm,1.19毫米,1.88 mm,RV为1.88 mm。然后,我们评估了ASSD不确定性措施如何识别不良细分的程度。为了利用ASSD不确定性测量,可以设置阈值,使得对于手动评审,标记了以上阈值的不确定性的任何分段。这希望导致数据集中减少的分割数量减少。图形预测_plots_assd显示了数据集中剩余的分段不良的图像的分数,并且当不确定性阈值变化时标记为手动校正的图像的分数,即(正面 - 真阳性)* vs *(真正的阳性+误报),其中代表不良分割。
As we decreased the uncertainty threshold (top left to bottom right in Figureprecision_plots_assd), we flagged more images for manual correction and the number of images with poor segmentation was decreased. The first point on the curves corresponds to a threshold where none of the images are reviewed while the last point indicates that all of the images are reviewed. This is similar to a receiver operating curve with the consideration that the total number of positives or images with poor predicted segmentation is different for each method. This allows for comparison between all the uncertainty estimation methods. Specifically, a curve closer to the bottom left corner or a curve with smaller area under the curve ( $ \mathrm{AUC} $ ) indicates that the algorithm provides better initial segmentation and/or its uncertainty is a good indicator of segmentation accuracy. Figure precision_plots_assd shows that all the methods performed similarly for detecting poor LV, Myo, and RV segmentation with Deep Ensembles having slightly lower $ \mathrm{AUC} $ than the other methods.

The thresholds of the uncertainty measures for flagging images for manual review can be adjusted based on the requirements of the application. This approach provides a way to identify images to review and may result in substantial time savings. For example, using the ASSD uncertainty measure for the Deep Ensembles method, 48%, 38%, and 31% of the images required manual review in order to reduce the number of images with poor LV, Myo, or RV segmentation to 5% of the test dataset, respectively (Figure precision_plots_assd). In contrast, without using uncertainty measures and assuming no other information about the images is used, approximately 75% of the images need to be reviewed to reduce the proportion of poor segmentation to 5%. Furthermore, using the ASSD uncertainty measure resulted in more time savings compared to a naive approach of reviewing images based on slice position. Specifically, since the segmentation is usually worse on basal and apical slices, a naive approach for segmentation quality control is to first review all of the most basal and most apical slices, followed by the second-most basal and second-most apical slices, and so on. We refer to this approach as using the slice position as a heuristic for segmentation quality control. As shown in Figure precision_plots_assd, using uncertainty measures is more advantageous than this naive approach as evidenced by a lower $ \mathrm{AUC} $ and a smaller number of images to review to have 5% poor segmentation remaining. Assuming a dataset of 10,000 subjects each with 10 slices and manual segmentation of 30 seconds per structure per slice , using the ASSD uncertainty measure results in $ \sim $ 940 hours of time savings compared to reviewing the images randomly, and $ \sim $ 580 hours of time savings compared to using the slice position heuristic to achieve 5% poor segmentation remaining.
随着我们减少不确定性阈值(图预先绘制的左上角到右下角),我们将更多的图像标记为手动校正,并且分段不良的图像数量减少。曲线上的第一点对应于阈值,其中缺任何图像,而最后一个点指示正在回顾所有图像。这类似于接收器操作曲线,考虑到预测分割的阳性或图像的总数对于每种方法不同。这允许在所有不确定性估计方法之间进行比较。具体地,更靠近左下角的曲线或曲线下方具有较小区域的曲线($\mathrm{AUC} $)表示该算法提供更好的初始分割和/或其不确定性是分割精度的良好指标。图Precision_plots_assd显示,所有方法都执行类似地用于检测差的LV,MyO和RV分段,与Duple Sensembles略低于$\mathrm{AUC} $的$\mathrm{AUC} $。
可以根据应用的要求调整标记用于手动审查的图像的不确定性措施的阈值。这种方法提供了一种识别图像以审查的方法,并且可能会节省大量时间。例如,使用ASSD不确定性测量的深度融合方法,48%,38%和31%的图像所需的手动审查,以减少LV,MyO或RV分段差的图像数量为5%分别测试数据集(图PROCISION_PLOTS_ASSD)。相反,在不使用不确定性测量和假设使用有关图像的其他信息的情况下,需要审查大约75%的图像,以将分段不良的比例降低到5%。此外,与基于切片位置审查图像的天真的方法相比,使用ASSD不确定性测量产生更多时间。具体地,由于分段通常在基础和顶端切片上更差,因此分割质量控制的天真方法是首先介绍所有最基的和最顶端的切片,然后是第二个最基的和第二个顶端切片,以及很快。我们将这种方法称为使用切片位置作为分割质量控制的启发式。如图所示,使用不确定性测量比这种天真的方法更有利,如较低的$\mathrm{AUC} $和较少数量的图像,以审查剩余5%的分段。假设10,000个科目的数据集每条有10个切片和手动分割,每片结构30秒,使用ASSD不确定性测量结果在$\sim $ 940小时的时间内节省,与随机审查图像,$\sim $ 580小时与使用切片位置启发式相比,节省时间储蓄仍然存在5%的细分。
Training Time
All experiments were performed on an Nvidia P100 GPU with 12GB of memory. MC Dropout and BBB required $ \sim $ 35 hours and $ \sim $ 47 hours for training, respectively. For these Bayesian methods, generating 50 prediction samples during testing required 1.5 seconds for each 2D image. The Deep Ensembles method consisted of 10 copies of the plain U-net and was trained in parallel separately. Each plain U-net required $ \sim $ 12 hours for training and $ \sim $ 0.1 seconds for prediction of a 2D image. Calculation of the predictive entropy, mutual information, Dice, and ASSD uncertainty measures required approximately 0.05, 0.15, 0.15, and 0.4 seconds per image, respectively.
训练时间
所有实验均在具有12GB的内存的NVIDIA P100 GPU上进行。 MC辍学和BBB所需$\sim $ 35小时和$\sim $ 47小时培训。对于这些贝叶斯方法,在测试期间在测试期间生成50个预测样本。每个2D图像需要1.5秒。深度合奏方法由普通U-Net的10个副本组成,并分别并行培训。每个普通U-Net所需的$\sim $ 12小时训练和$\sim $ 0.1秒用于预测2D图像。计算预测熵,互信息,骰子和ASSD不确定性测量的每张图像的约0.05,0.15,0.15和0.4秒。
Discussion
讨论
BBB \emph MC Dropout \emph Deep Ensemble
In this work, we evaluated and compared different Bayesian and non-Bayesian approaches for estimating uncertainty in neural networks for cardiac MRI segmentation. Here, we discuss similarities and differences about how uncertainty is learned in these methods, what was learned after training, and relate these differences to the quality of the predictive uncertainties on out-of-distribution images. While uncertainty in neural network parameters is learned automatically in BBB, this can be tuned by changing the dropout rate in MC Dropout. For the UKBB dataset, a small dropout rate of 0.1 in the middle layers performed better than other MC Dropout models in terms of segmentation accuracy and probability calibration. This is different from other studies which commonly used a dropout rate of 0.5 and may be because of the large amount of relatively uniform training data (i.e., images acquired following the same MR protocol and labelled following the same guidelines). The dropout rate hyperparameters for MC Dropout obtained through grid search correspond to the weight uncertainties learned by BBB to some extent. Figure weight_stats shows that the standard deviation of the weights learned by BBB is lower in the early layers and higher in the middle layers of the U-net. This is similar to MC Dropout with no dropout in the early layers and some dropout on the middle layers. Having greater dropout rate in middle layers is common in other studies .

The effect of dropout rate in MC Dropout is shown in the experiments with dataset shift (Figure metrics_correlations, left). A dropout rate of 0.1 yielded higher segmentation accuracy but lower uncertainty whereas a dropout rate of 0.5 resulted in lower segmentation accuracy but higher predictive uncertainty. BBB was able to mitigate this issue by learning a mean and standard deviation for each weight, resulting in comparable or higher segmentation accuracy with moderate predictive uncertainties between MCD-0.
bbb \ emph mc agropout \ emph深组合
在这项工作中,我们评估并比较了不同的贝叶斯和非贝叶斯的方法,以估算心脏MRI分割神经网络的不确定性。在这里,我们讨论了对这些方法中学到的不确定性的相似性和差异,培训后如何学到,并将这些差异与预测性不确定性的质量相关联而言。虽然在BBB中自动学习神经网络参数的不确定性,但可以通过更改MC丢失中的丢失率来调整这一点。对于UKBB数据集,中间层中的小幅放大速率比分段精度和概率校准在其他MC辍学模型中更好地执行。这与其他研究不同,这些研究通常使用0.5的辍学率,并且可能是由于大量相对均匀的训练数据(即,在相同的MR协议上获取的图像并按照相同的指导方式标记)。通过网格搜索获得的MC辍学的辍学率超级参数对应于BBB在一定程度上学习的权重不确定性。图重量_STATS表明,BBB学习的权重的标准偏差在U-Net的中间层中较低,更高。这类似于MC辍学,早期没有辍学,中间层上的一些丢失。在其他研究中具有更大的中层辍学率是常见的。
在数据集移位的实验中显示了MC辍学中的辍学率的效果(图METRICS_CORRELIANCS,左侧)。 0.1的辍学率产生较高的分割精度,但不确定性较低,而辍学率为0.5导致降低的分割精度,但更高的预测性不确定性。 BBB能够通过学习每种重量的平均值和标准偏差来减轻这个问题,导致在MCD-0之间具有适度的预测性不确定性的相当或更高的分割精度。
1 and MCD-0.5 (Figure metrics_correlations, left). In Deep Ensembles, the uncertainty in the weights is not learned or tuned but instead stems from the random initialization of the weights and stochasticity of the training procedure. Each model in the ensemble would then learn a local minimum which are combined to form a prediction and uncertainty estimate. This was found to outperform BBB in terms of segmentation accuracy and probability calibration in most cases. This may be because the approximate posterior learned by BBB covered only one (or a few) mode(s) of the true posterior and a small neighbourhood around each mode, as opposed to multiple local modes in Deep Ensembles. Based on our observations, the histogram of the weights learned by each member in Deep Ensembles was very similar to each other. Techniques such as canonical correlation analysis can be used to compare the representations (i.e., feature maps) between different members of the ensemble and to better understand model uncertainty . While Deep Ensembles performed similarly or better than BBB on images with blurring or stretching distortions, BBB outperformed Deep Ensembles on images with noise distortions. A difference in these distortions is that the noise was added independently per pixel while blurring and stretching were applied for each structure as a whole. This suggests that BBB is more suitable for capturing uncertainty in low-level pixelwise variations while Deep Ensemble is better for modelling uncertainty in structural variations. Bayesian approaches and Deep Ensembles can be used to improve segmentation accuracy on datasets which are slightly shifted compared to the training dataset. While the segmentation accuracies for testing on the ACDC dataset using the models trained on UKBB dataset are lower than segmentation accuracies obtained using models trained and tested both on the ACDC dataset , the algorithms employed in this work may be combined with other techniques that are specifically designed to solve this problem, e.g., style augmentation or domain adversarial training .
1和MCD-0.5(图METRICS_CORRELATIONS,左侧)。在深度集合中,不学习或调整权重中的不确定性,而是从训练过程的重量和随机性的随机初始化源。然后,集合中的每个模型将学习局部最小值,该局部最小是形成预测和不确定性估计的。在大多数情况下,发现在分割精度和概率校准方面以BBB优于BBB。这可能是因为BBB所学到的近似后后部只覆盖了每种模式的每种模式的真实后的一个(或几个)模式,而不是深度集合中的多种本地模式。基于我们的观察,深融合中每个成员学习的重量的直方图彼此非常相似。诸如规范相关分析的技术可用于比较集合的不同成员之间的表示(即特征映射)并更好地理解模型不确定性。虽然深度集合在具有模糊或拉伸扭曲的图像上类似或更好地进行或更好地进行,但BBB在具有噪声扭曲的图像上表现出深度的深度集成。这些失真的差异是每像素独立地添加噪声,而整个结构施加模糊和拉伸。这表明BBB更适合于捕获低级像素变化中的不确定性,而深度集合更好地用于在结构变化中建模不确定性。贝叶斯方法和深度集合可用于提高数据集的分段精度,与训练数据集相比略微移位。虽然使用在UKBB数据集上培训的模型对ACDC数据集进行测试的分割精度低于使用培训和在ACDC数据集上进行的模型获得的分段精度,但该工作中采用的算法可以与专门设计的其他技术相结合解决这个问题,例如,风格增强或域对抗训练。
Pixelwise and Structural Uncertainty Measures
We introduced pixelwise and structural uncertainty measures to quantify the predictive uncertainty, and demonstrated the utility of these metrics for segmentation quality control. Both pixelwise and structural uncertainty measures can be used depending on the application. As the segmentation problem was formulated as pixel classification, pixelwise uncertainty measures are straightforward to obtain. These allow users to visualize which pixels and which areas are potentially problematic (Figure noise_distortion_image). However, segmentation is often performed at the image-level and slice by slice. Therefore, in developing uncertainty measures for determining problematic segmentation, an image-level uncertainty measure is required. Accordingly, we showed that structural uncertainty measures are correlated with segmentation accuracy metrics such as ASSD. It is important to note that the predictive uncertainty measures reflect the neural network uncertainty, which is different from human uncertainty. An example is the image with heavy noise in the last row in Figure noise_distortion_image. It is expected that human observers can manually segment this image with low observer variability; however, since the image is very different from the training data, the neural networks were not able to generate a reasonable segmentation and showed high predictive entropy and mutual information for the entire cardiac structure.
Pixelwise和结构不确定度测量
我们介绍了像素向上和结构性不确定性措施,以量化预测性的不确定性,并证明了这些度量的效用进行了分割质量控制。可以根据应用使用PixElWive和结构不确定性措施。随着分段问题被制定为像素分类,Pix透明的不确定度措施很简单地获得。这些允许用户可视化哪些像素以及哪个区域可能存在问题(图诊断_distorgatorrage_Image)。但是,分段通常在图像级和切片上执行切片。因此,在开发用于确定有问题分割的不确定性措施时,需要一种图像级不确定性度量。因此,我们表明,结构不确定性措施与分割精度指标相关,例如ASSD。值得注意的是,预测性不确定性措施反映了神经网络不确定性,这与人类不确定性不同。一个例子是图中的噪声噪声中的噪声噪声_DISTORGATORGAGE_IMAGE中的图像。预计人类观察者可以用低观察者变异性地手动分割此图像;然而,由于图像与训练数据非常不同,所以神经网络无法产生合理的分割,并且为整个心脏结构显示了高预测熵和相互信息。
Segmentation Quality Control
We showed that the uncertainty measures have decent to good correlations with segmentation accuracy. This could have been negatively affected by noisy ground truth segmentation. Framing segmentation quality control as a binary classification problem instead of evaluating the correlations or predicting the segmentation accuracy metrics alleviates the problem of noise in ground truth segmentation. In this regard, we defined poor segmentation using a threshold on the ASSD between the predicted and ground truth segmentation. This definition of a poor segmentation was adopted based on discussions with clinical collaborators; however, it can be modified depending on the application. For example, other segmentation accuracy metrics such as Hausdorff distance or misclassification area can be used and the framework for evaluating uncertainty measures described in this work can be applied directly. Other studies in segmentation quality control include directly predicting segmentation accuracy or comparing the predicted segmentation to a reference database. trained a random forest classifier to predict a binary label of correct or incorrect segmentation. The examples of correct segmentation were generated using the ground truth segmentations while the incorrect segmentations were generated by deforming or translating the ground truth segmentations. used a 3D residual network to directly predict the Dice score of a segmentation from an image-segmentation pair. The network was trained and tested on a dataset created using a random forest segmentation algorithm. These classifiers are agnostic to how the segmentation was generated.
分割质量控制
我们认为不确定性措施具有与分割精度的良好相关性。这可能对嘈杂的地面真相细分产生负面影响。框架分割质量控制作为二进制分类问题,而不是评估相关性或预测分割精度度量,减轻了地面真实分割中的噪声问题。在这方面,我们使用预测和地面事实分割之间的ASSD的阈值来定义差的分割。基于临床合作者的讨论,采用了这种不良分割的定义;但是,它可以根据应用程序修改。例如,可以使用其他分割精度度量,例如Hausdorff距离或错误分类区域,并且可以直接应用用于评估本工作中描述的不确定性措施的框架。分割质量控制的其他研究包括直接预测分割精度或将预测的分段与参考数据库进行比较。培训了随机林分类器,以预测正确或不正确的分割的二进制标签。使用地面真理分段生成正确分割的示例,而通过使地面真相分段变形或转换地产生不正确的分割。使用3D残余网络直接从图像分割对中预测分割的骰子分数。在使用随机林分段算法创建的数据集上培训并测试网络。这些分类器对于如何生成分割是不可知的。
However, these approaches are dependent on training with some expected segmentation failures, which may be challenging to incorporate during training. In contrast, our approach used uncertainty measures to detect poor segmentation and is explainable. Instead of using a learning algorithm to determine segmentation quality, we used model uncertainty which emerges intrinsically when training the segmentation algorithm. A slight limitation of our approach is that sampling during the testing phase required up to 50x more computation time compared to a single prediction but this may be accelerated through parallelization. Another approach for segmentation quality control is using a modified version of Reverse Classification Accuracy to predict the segmentation quality of an image-segmentation pair . This approach requires a reference database of images with ground truth segmentation. Each reference image is registered to the test image and the associated reference segmentations are warped accordingly. The warped reference segmentations are considered as potential segmentations of the test image. The quality of the test image segmentation is estimated by comparing the potential segmentations and test segmentation. A limitation of this approach is that it requires a long time to generate the segmentation quality prediction, mainly due to the registration steps.
然而,这些方法取决于培训,一些预期的细分失败,这可能在培训期间纳入挑战。相比之下,我们的方法使用不确定性措施来检测不良分割,可扩展理。而不是使用学习算法来确定分割质量,而是使用模型不确定性,在训练分割算法时本质上出现。对我们的方法的略有限制是测试阶段在测试阶段期间采样高达50倍的计算时间与单个预测相比,但这可以通过并行化加速。用于分割质量控制的另一种方法是使用反向分类精度的修改版本来预测图像分割对的分割质量。该方法需要具有地面真实分割的图像的参考数据库。每个参考图像被登记到测试图像,并且相应地扭转相关的参考分割。翘曲的参考分割被认为是测试图像的潜在分割。通过比较潜在的分段和测试分割来估计测试图像分割的质量。这种方法的限制是它需要很长时间来产生分割质量预测,主要原因是注册步骤。
Conclusions
In this work, we compared Bayesian and non-Bayesian methods, namely BBB, MC Dropout, and Deep Ensembles for cardiac MRI segmentation and showed the utility of these algorithms for segmentation quality control. We found that Deep Ensembles performed better in terms of segmentation accuracy and probability calibration on in-distribution and out-of-distribution datasets. However, BBB outperformed Deep Ensembles on images with noise distortions. Additionally, we identified a limitation of current methods in terms of predictive uncertainty on images with heavy distortions. Furthermore, we showed that the ASSD uncertainty measure correlated well with the ASSD segmentation accuracy on both in-distribution and out-of-distribution datasets. For the task of segmentation quality control, ASSD from all methods are equally useful in predicting problematic segmentations. Using uncertainty measures can result in substantial time savings by reducing the number of images that needs to be reviewed. We acknowledge the use of the facilities of Compute Canada. This work was funded by Canadian Institutes of Health Research (CIHR) MOP: #93531, Ontario Research Fund and GE Healthcare. FG is supported by a Banting postdoctoral fellowship. This work was partly funded by the European Union’s Horizon 2020 research and innovation programme under grant agreement No 825903 (euCanSHare project). SEP acts as a paid consultant to Circle Cardiovascular Imaging Inc., Calgary, Canada and Servier. SEP acknowledges support from the National Institute for Health Research (NIHR) Biomedical Research Centre at Barts, from the SmartHeart EPSRC programme grant (EP/P001009/1) and the London Medical Imaging and AI Centre for Value-Based Healthcare. SEP acknowledges support from the CAP-AI programme, London’s first AI enabling programme focused on stimulating growth in the capital’s AI sector. SEP, SN and SKP acknowledge the British Heart Foundation for funding the manual analysis to create a cardiovascular magnetic resonance imaging reference standard for the UK Biobank imaging resource in 5000 CMR scans (PG/14/89/31194). This project was enabled through access to the Medical Research Council eMedLab Medical Bioinformatics infrastructure, supported by the Medical Research Council (MR/L016311/1). This research was conducted using the UK Biobank resource under Application 2964. UK Biobank will make the data available to all bona fide researchers for all types of health-related research that is in the public interest, without preferential or exclusive access for any person. All researchers will be subject to the same application process and approval criteria as specified by UK Biobank. For the detailed access procedure see http://www.ukbiobank.ac.uk/register-apply/. Please see the supplementary material for detailed tables of results.
结论
在这项工作中,我们比较了贝叶斯和非贝叶斯方法,即BBB,MC辍学和心脏MRI分割的深度集成,并显示了这些算法进行分割质量控制的效用。我们发现,在分配准确性和分配方式和分发外部数据集的概率校准方面,深度集合更好。然而,BBB在具有噪声扭曲的图像上表现出深入的集成。另外,我们确定了对具有严重扭曲图像的预测性不确定性的当前方法的限制。此外,我们表明,ASSD不确定性测量与分布式和分配外数据集的ASSD分段精度相比好。对于分割质量控制的任务,来自所有方法的ASSD在预测有问题的细分方面同样有用。使用不确定性措施可以通过减少需要审查的图像数量来产生大量的节省时间。我们承认使用加拿大计算设施。
这项工作由加拿大卫生研究院资助(CIHR)MOP:\#93531,安大略研究基金和GE Healthcare。 FG由Banting Proddoctoral奖学金支持。这项工作部分由欧洲联盟的地平线2020的研究和创新计划部分资助,根据拨款协议没有825903(宇航员项目)。 SEP充当付费顾问来圈出心血管成像Inc.,卡尔加里,加拿大和服务器。 SEP认识到巴斯特国家卫生研究所(NIHR)生物医学研究中心的支持,从SMARTHEART EPSRC计划授予(EP / P001009 / 1)和伦敦医学成像和基于价值的医疗保健中心。 SEP认识到CAP-AI计划的支持,伦敦的第一个AI支持计划专注于刺激首都AI部门的增长。 SEP,SN和SKP承认英国心脏基金会为5000厘米扫描(PG / 14/89/31194)中英国Biobank成像资源创造心血管磁共振成像参考标准。该项目通过访问医学研究委员会Emedlab医学生物信息学基础设施,支持该项目,由医学研究委员会支持(MR / L016311 / 1)。该研究是在申请2964的申请中使用英国Biobank资源进行的。英国Biobank将使所有与公共利益的所有类型的健康相关研究中的所有类型的Bona Fife研究人员提供数据,而不优惠或独家访问。所有研究人员都将受到英国Biobank指定的相同申请流程和批准标准。有关详细访问过程,请参阅http://www.ukbiobank.ac.uk/register-apply/。有关详细的结果,请参阅辅助材料。
